The hantavirus nucleocapsid protein recognizes specific features of the viral RNA panhandle and is altered in conformation upon RNA binding
- PMID: 15650206
- PMCID: PMC544099
- DOI: 10.1128/JVI.79.3.1824-1835.2005
The hantavirus nucleocapsid protein recognizes specific features of the viral RNA panhandle and is altered in conformation upon RNA binding
Abstract
Hantaviruses are tripartite negative-sense RNA viruses and members of the Bunyaviridae family. The nucleocapsid (N) protein is the principal structural component of the viral capsid. N forms a stable trimer that specifically recognizes the panhandle structure formed by the viral RNA termini. We used trimeric glutathione S-transferase (GST)-N protein and small RNA panhandles to examine the requirements for specific recognition by Sin Nombre hantavirus N. Trimeric GST-N recognizes the panhandles of the three viral RNAs (S, M, and L) with high affinity, whereas the corresponding plus-strand panhandles of the complementary RNA are recognized with lower affinity. Based on analysis of nucleotide substitutions that alter either the higher-order structure of the panhandle or the primary sequence of the panhandle, both secondary structure and primary sequence are necessary for stable interaction with N. A panhandle 23 nucleotides long is necessary and sufficient for high-affinity binding by N, and stoichiometry calculations indicate that a single N trimer interacts with a single panhandle. Surprisingly, displacement of the panhandle structure away from the terminus does not eliminate recognition by N. The binding of N to the panhandle is an entropy-driven process resulting in initial stable N-RNA interaction followed by a conformational change in N. Taken together, these data provide insight into the molecular events that take place during interaction of N with the panhandle and suggest that specific high-affinity interaction between an RNA binding domain of trimeric N and the panhandle is required for encapsidation of the three viral RNAs.
Figures





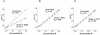
Similar articles
-
Hantavirus N protein exhibits genus-specific recognition of the viral RNA panhandle.J Virol. 2006 Nov;80(22):11283-92. doi: 10.1128/JVI.00820-06. Epub 2006 Sep 13. J Virol. 2006. PMID: 16971445 Free PMC article.
-
Trimeric hantavirus nucleocapsid protein binds specifically to the viral RNA panhandle.J Virol. 2004 Aug;78(15):8281-8. doi: 10.1128/JVI.78.15.8281-8288.2004. J Virol. 2004. PMID: 15254200 Free PMC article.
-
Identification of a region of hantavirus nucleocapsid protein required for RNA chaperone activity.RNA Biol. 2010 Nov-Dec;7(6):830-7. doi: 10.4161/rna.7.6.13862. Epub 2010 Nov 1. RNA Biol. 2010. PMID: 21378500 Free PMC article.
-
The Nucleocapsid of Paramyxoviruses: Structure and Function of an Encapsidated Template.Viruses. 2021 Dec 9;13(12):2465. doi: 10.3390/v13122465. Viruses. 2021. PMID: 34960734 Free PMC article. Review.
-
The nucleocapsid of vesicular stomatitis virus.Sci China Life Sci. 2012 Apr;55(4):291-300. doi: 10.1007/s11427-012-4307-x. Epub 2012 May 9. Sci China Life Sci. 2012. PMID: 22566085 Review.
Cited by
-
RNA Encapsidation and Packaging in the Phleboviruses.Viruses. 2016 Jul 15;8(7):194. doi: 10.3390/v8070194. Viruses. 2016. PMID: 27428993 Free PMC article. Review.
-
The N terminus of Rift Valley fever virus nucleoprotein is essential for dimerization.J Virol. 2005 Sep;79(18):11974-80. doi: 10.1128/JVI.79.18.11974-11980.2005. J Virol. 2005. PMID: 16140773 Free PMC article.
-
Hantavirus N protein exhibits genus-specific recognition of the viral RNA panhandle.J Virol. 2006 Nov;80(22):11283-92. doi: 10.1128/JVI.00820-06. Epub 2006 Sep 13. J Virol. 2006. PMID: 16971445 Free PMC article.
-
Crystal Structure of the Core Region of Hantavirus Nucleocapsid Protein Reveals the Mechanism for Ribonucleoprotein Complex Formation.J Virol. 2015 Nov 11;90(2):1048-61. doi: 10.1128/JVI.02523-15. Print 2016 Jan 15. J Virol. 2015. PMID: 26559827 Free PMC article.
-
Retrospective on the all-in-one retroviral nucleocapsid protein.Virus Res. 2014 Nov 26;193:2-15. doi: 10.1016/j.virusres.2014.05.011. Epub 2014 Jun 4. Virus Res. 2014. PMID: 24907482 Free PMC article. Review.
References
-
- Bougie, I., and M. Bisaillon. 2003. Initial binding of the broad spectrum antiviral nucleoside ribavirin to the hepatitis C virus RNA polymerase. J. Biol. Chem. 278:52471-52478. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials

