Anti-CXCR4 monoclonal antibodies recognizing overlapping epitopes differ significantly in their ability to inhibit entry of human immunodeficiency virus type 1
- PMID: 15650218
- PMCID: PMC544137
- DOI: 10.1128/JVI.79.3.1930-1933.2005
Anti-CXCR4 monoclonal antibodies recognizing overlapping epitopes differ significantly in their ability to inhibit entry of human immunodeficiency virus type 1
Abstract
CXCR4 is one of two physiologically relevant human immunodeficiency type 1 (HIV-1) entry coreceptors. Studies of CXCR4 mutants have not clearly identified the determinants of coreceptor function and specificity. We therefore used a panel of monoclonal antibodies to further elucidate CXCR4 expression, structure, and function. Our findings show the existence of conformational subpopulations of CXCR4 that are in equilibrium on the cell surface but are not cell type specific as previously reported. HIV-1 X4 isolates can interact with multiple CXCR4 conformations in order to gain entry into target cells.
Figures

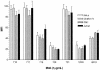
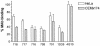

Similar articles
-
Unique monoclonal antibody recognizing the third extracellular loop of CXCR4 induces lymphocyte agglutination and enhances human immunodeficiency virus type 1-mediated syncytium formation and productive infection.J Virol. 2001 Dec;75(23):11534-43. doi: 10.1128/JVI.75.23.11534-11543.2001. J Virol. 2001. PMID: 11689635 Free PMC article.
-
Development of a cell-based enzyme-linked immunosorbent assay for high-throughput screening of HIV type 1 entry inhibitors targeting the coreceptor CXCR4.AIDS Res Hum Retroviruses. 2003 Nov;19(11):947-55. doi: 10.1089/088922203322588297. AIDS Res Hum Retroviruses. 2003. PMID: 14678601
-
Antigenically distinct conformations of CXCR4.J Virol. 2001 Oct;75(19):8957-67. doi: 10.1128/JVI.75.19.8957-8967.2001. J Virol. 2001. PMID: 11533159 Free PMC article.
-
Cyclic zinc-dithiocarbamate-S,S'-dioxide blocks CXCR4-mediated HIV-1 infection.Biochem Biophys Res Commun. 2000 Jun 7;272(2):351-6. doi: 10.1006/bbrc.2000.2779. Biochem Biophys Res Commun. 2000. PMID: 10833417
-
Cryptic nature of a conserved, CD4-inducible V3 loop neutralization epitope in the native envelope glycoprotein oligomer of CCR5-restricted, but not CXCR4-using, primary human immunodeficiency virus type 1 strains.J Virol. 2005 Jun;79(11):6957-68. doi: 10.1128/JVI.79.11.6957-6968.2005. J Virol. 2005. PMID: 15890935 Free PMC article.
Cited by
-
Discovery of Bis-Imidazoline Derivatives as New CXCR4 Ligands.Molecules. 2023 Jan 24;28(3):1156. doi: 10.3390/molecules28031156. Molecules. 2023. PMID: 36770826 Free PMC article.
-
Mapping Interaction Sites on Human Chemokine Receptors by Deep Mutational Scanning.J Immunol. 2018 Jun 1;200(11):3825-3839. doi: 10.4049/jimmunol.1800343. Epub 2018 Apr 20. J Immunol. 2018. PMID: 29678950 Free PMC article.
-
Detecting Cell Surface Expression of the G Protein-Coupled Receptor CXCR4.Methods Mol Biol. 2018;1722:151-164. doi: 10.1007/978-1-4939-7553-2_10. Methods Mol Biol. 2018. PMID: 29264804 Free PMC article.
-
Identifying and Assessing Putative Allosteric Sites and Modulators for CXCR4 Predicted through Network Modeling and Site Identification by Ligand Competitive Saturation.J Phys Chem B. 2024 May 30;128(21):5157-5174. doi: 10.1021/acs.jpcb.4c00925. Epub 2024 Apr 22. J Phys Chem B. 2024. PMID: 38647430 Free PMC article.
-
Chemokine receptor-specific antibodies in cancer immunotherapy: achievements and challenges.Front Immunol. 2015 Jan 30;6:12. doi: 10.3389/fimmu.2015.00012. eCollection 2015. Front Immunol. 2015. PMID: 25688243 Free PMC article. Review.
References
-
- Berger, E. A., P. M. Murphy, and J. M. Farber. 1999. Chemokine receptors as HIV-1 coreceptors: roles in viral entry, tropism, and disease. Annu. Rev. Immunol. 17:657-700. - PubMed
-
- Brelot, A., N. Heveker, M. Montes, and M. Alizon. 2000. Identification of residues of CXCR4 critical for human immunodeficiency virus coreceptor and chemokine receptor activities. J. Biol. Chem. 275:23736-23744. - PubMed
-
- Chabot, D. J., P. F. Zhang, G. V. Quinnan, and C. C. Broder. 1999. Mutagenesis of CXCR4 identifies important domains for human immunodeficiency virus type 1 X4 isolate envelope-mediated membrane fusion and virus entry and reveals cryptic coreceptor activity for R5 isolates. J. Virol. 73:6598-6609. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

