Evolution of the isoprene biosynthetic pathway in kudzu
- PMID: 15653811
- PMCID: PMC1065370
- DOI: 10.1104/pp.104.054445
Evolution of the isoprene biosynthetic pathway in kudzu
Abstract
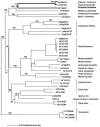
Isoprene synthase converts dimethylallyl diphosphate, derived from the methylerythritol 4-phosphate (MEP) pathway, to isoprene. Isoprene is made by some plants in substantial amounts, which affects atmospheric chemistry, while other plants make no isoprene. As part of our long-term study of isoprene synthesis, the genetics of the isoprene biosynthetic pathway of the isoprene emitter, kudzu (Pueraria montana), was compared with similar genes in Arabidopsis (Arabidopsis thaliana), which does not make isoprene. The MEP pathway genes in kudzu were similar to the corresponding Arabidopsis genes. Isoprene synthase genes of kudzu and aspen (Populus tremuloides) were cloned to compare their divergence with the divergence seen in MEP pathway genes. Phylogenetic analysis of the terpene synthase gene family indicated that isoprene synthases are either within the monoterpene synthase clade or sister to it. In Arabidopsis, the gene most similar to isoprene synthase is a myrcene/ocimene (acyclic monoterpenes) synthase. Two phenylalanine residues found exclusively in isoprene synthases make the active site smaller than other terpene synthase enzymes, possibly conferring specificity for the five-carbon substrate rather than precursors of the larger isoprenoids. Expression of the kudzu isoprene synthase gene in Arabidopsis caused Arabidopsis to emit isoprene, indicating that whether or not a plant emits isoprene depends on whether or not it has a terpene synthase capable of using dimethylallyl diphosphate.
Figures









Similar articles
-
In Planta Recapitulation of Isoprene Synthase Evolution from Ocimene Synthases.Mol Biol Evol. 2017 Oct 1;34(10):2583-2599. doi: 10.1093/molbev/msx178. Mol Biol Evol. 2017. PMID: 28637270 Free PMC article.
-
Rapid regulation of the methylerythritol 4-phosphate pathway during isoprene synthesis.Plant Physiol. 2004 Aug;135(4):1939-45. doi: 10.1104/pp.104.043737. Epub 2004 Jul 30. Plant Physiol. 2004. PMID: 15286290 Free PMC article.
-
Biosynthesis of isoprene in Escherichia coli via methylerythritol phosphate (MEP) pathway.Appl Microbiol Biotechnol. 2011 Jun;90(6):1915-22. doi: 10.1007/s00253-011-3199-1. Epub 2011 Apr 6. Appl Microbiol Biotechnol. 2011. PMID: 21468716
-
[Advances in isoprene synthase research].Sheng Wu Gong Cheng Xue Bao. 2017 Nov 25;33(11):1802-1813. doi: 10.13345/j.cjb.170040. Sheng Wu Gong Cheng Xue Bao. 2017. PMID: 29202517 Review. Chinese.
-
An account of cloned genes of Methyl-erythritol-4-phosphate pathway of isoprenoid biosynthesis in plants.Curr Issues Mol Biol. 2009;11 Suppl 1:i35-45. Epub 2009 Feb 2. Curr Issues Mol Biol. 2009. PMID: 19193963 Review.
Cited by
-
Identification of a fungal 1,8-cineole synthase from Hypoxylon sp. with specificity determinants in common with the plant synthases.J Biol Chem. 2015 Mar 27;290(13):8511-26. doi: 10.1074/jbc.M114.636159. Epub 2015 Feb 3. J Biol Chem. 2015. PMID: 25648891 Free PMC article.
-
Domestication and the storage starch biosynthesis pathway: signatures of selection from a whole sorghum genome sequencing strategy.Plant Biotechnol J. 2016 Dec;14(12):2240-2253. doi: 10.1111/pbi.12578. Epub 2016 Jun 11. Plant Biotechnol J. 2016. PMID: 27155090 Free PMC article.
-
Prediction of function for the polyprenyl transferase subgroup in the isoprenoid synthase superfamily.Proc Natl Acad Sci U S A. 2013 Mar 26;110(13):E1196-202. doi: 10.1073/pnas.1300632110. Epub 2013 Mar 14. Proc Natl Acad Sci U S A. 2013. PMID: 23493556 Free PMC article.
-
Evidence that light, carbon dioxide, and oxygen dependencies of leaf isoprene emission are driven by energy status in hybrid aspen.Plant Physiol. 2009 Sep;151(1):448-60. doi: 10.1104/pp.109.141978. Epub 2009 Jul 8. Plant Physiol. 2009. PMID: 19587097 Free PMC article.
-
Biosynthesis, accumulation and emission of carotenoids, alpha-tocopherol, plastoquinone, and isoprene in leaves under high photosynthetic irradiance.Photosynth Res. 2007 May;92(2):163-79. doi: 10.1007/s11120-007-9204-y. Epub 2007 Jul 17. Photosynth Res. 2007. PMID: 17634750 Review.
References
-
- Altschul SF, Gish W, Miller W, Meyers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215: 403–410 - PubMed
-
- Aubourg S, Lecharny A, Bohlmann J (2002) Genomic analysis of the terpenoid synthase (AtTPS) gene family of Arabidopsis thaliana. Mol Genet Genomics 267: 730–745 - PubMed
-
- Bohlmann J, Martin D, Oldham NJ, Gershenzon J (2000) Terpenoid secondary metabolism in Arabidopsis thaliana: cDNA cloning, characterization, and functional expression of a myrcene/(E)-beta-ocimene synthase. Arch Biochem Biophys 375: 261–269 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

