Insertion/deletion and nucleotide polymorphism data reveal constraints in Drosophila melanogaster introns and intergenic regions
- PMID: 15654088
- PMCID: PMC1449560
- DOI: 10.1534/genetics.104.037689
Insertion/deletion and nucleotide polymorphism data reveal constraints in Drosophila melanogaster introns and intergenic regions
Abstract
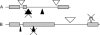
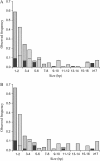
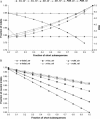
Our study of nucleotide sequence and insertion/deletion polymorphism in Drosophila melanogaster noncoding DNA provides evidence for selective pressures in both intergenic regions and introns (of the large size class). Intronic and intergenic sequences show a similar polymorphic deletion bias. Insertions have smaller sizes and higher frequencies than deletions, supporting the hypothesis that insertions are selected to compensate for the loss of DNA caused by deletion bias. Analysis of a simple model of selective constraints suggests that the blocks of functional elements located in intergenic sequences are on average larger than those in introns, while the length distribution of relatively unconstrained sequences interspaced between these blocks is similar in intronic and intergenic regions.
Figures
Similar articles
-
Analysis of conserved noncoding DNA in Drosophila reveals similar constraints in intergenic and intronic sequences.Genome Res. 2001 Aug;11(8):1335-45. doi: 10.1101/gr.178701. Genome Res. 2001. PMID: 11483574
-
Patterns of polymorphism and divergence from noncoding sequences of Drosophila melanogaster and D. simulans: evidence for nonequilibrium processes.Mol Biol Evol. 2005 Jan;22(1):51-62. doi: 10.1093/molbev/msh269. Epub 2004 Sep 29. Mol Biol Evol. 2005. PMID: 15456897 Review.
-
Rapid sequence turnover at an intergenic locus in Drosophila.Mol Biol Evol. 2004 Apr;21(4):670-80. doi: 10.1093/molbev/msh060. Epub 2004 Jan 22. Mol Biol Evol. 2004. PMID: 14739245
-
Intron length evolution in Drosophila.Mol Biol Evol. 2006 Nov;23(11):2203-13. doi: 10.1093/molbev/msl094. Epub 2006 Aug 21. Mol Biol Evol. 2006. PMID: 16923822
-
[Functional genomics in Drosophila melanogaster].Tanpakushitsu Kakusan Koso. 2001 Dec;46(16 Suppl):2436-40. Tanpakushitsu Kakusan Koso. 2001. PMID: 11802407 Review. Japanese. No abstract available.
Cited by
-
Strong mutational bias toward deletions in the Drosophila melanogaster genome is compensated by selection.Genome Biol Evol. 2013;5(3):514-24. doi: 10.1093/gbe/evt021. Genome Biol Evol. 2013. PMID: 23395983 Free PMC article.
-
Abundance and distribution of transposable elements in two Drosophila QTL mapping resources.Mol Biol Evol. 2013 Oct;30(10):2311-27. doi: 10.1093/molbev/mst129. Epub 2013 Jul 24. Mol Biol Evol. 2013. PMID: 23883524 Free PMC article.
-
Ubiquitous selective constraints in the Drosophila genome revealed by a genome-wide interspecies comparison.Genome Res. 2006 Jul;16(7):875-84. doi: 10.1101/gr.5022906. Epub 2006 Jun 2. Genome Res. 2006. PMID: 16751341 Free PMC article.
-
Multilocus phylogeography and phylogenetics using sequence-based markers.Genetica. 2009 Apr;135(3):439-55. doi: 10.1007/s10709-008-9293-3. Epub 2008 Jul 24. Genetica. 2009. PMID: 18651229 Review.
-
Contrasting patterns of sequence divergence and base composition between Drosophila introns and intergenic regions.Biol Lett. 2006 Dec 22;2(4):604-7. doi: 10.1098/rsbl.2006.0521. Biol Lett. 2006. PMID: 17148300 Free PMC article.
References
-
- Bergman, C. M., and M. Kreitman, 2001. Analysis of conserved noncoding DNA in Drosophila reveals similar constraints in intergenic and intronic sequences. Genome Res. 11: 1335–1345. - PubMed
-
- Blumenstiel, J. P., D. L. Hartl and E. R. Lozowsky, 2002. Patterns of insertion and deletion in contrasting chromatin domains. Mol. Biol. Evol. 19: 2211–2225. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

