The CYP2B2 phenobarbital response unit contains binding sites for hepatocyte nuclear factor 4, PBX-PREP1, the thyroid hormone receptor beta and the liver X receptor
- PMID: 15656786
- PMCID: PMC1138947
- DOI: 10.1042/BJ20041556
The CYP2B2 phenobarbital response unit contains binding sites for hepatocyte nuclear factor 4, PBX-PREP1, the thyroid hormone receptor beta and the liver X receptor
Abstract
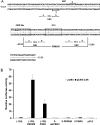
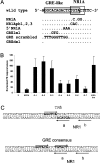
A 163 bp enhancer in the CYP2B2 5' flank confers PB (phenobarbital) inducibility and constitutes a PBRU (PB response unit). The PBRU contains several transcription factor binding sites, including NR1, NR2 and NR3, which are direct repeats separated by 4 bp of the nuclear receptor consensus half-site AGGTCA, as well as an ER (everted repeat) separated by 7 bp (ER-7). Constitutive androstane receptor (CAR)-RXR (retinoic X receptor) heterodimers are known to bind to NR1, NR2 and NR3. Electrophoretic mobility-shift analysis using nuclear extracts from livers of untreated or PB-treated rats revealed binding of several other proteins to different PBRU elements. Using supershift analysis and in vitro coupled transcription and translation, the proteins present in four retarded complexes were identified as TRbeta (thyroid hormone receptor beta), LXR (liver X receptor), HNF-4 (hepatocyte nuclear factor 4) and heterodimers of PBX-PREP1 (pre-B cell homoeobox-Pbx regulatory protein 1). LXR-RXR heterodimers bound to NR3 and TRbeta bound to NR3, NR1 and ER-7, whereas the PBX-PREP1 site is contained within NR2. The HNF-4 site overlaps with NR1. A mutation described previously, GRE1m1, which decreases PB responsiveness, increased the affinity of this site for HNF-4. The PBRU also contains a site for nuclear factor 1. The PBRU thus contains a plethora of transcription factor binding sites. The profiles of transcription factor binding to NR1 and NR3 were quite similar, although strikingly different from, and more complex than, that of NR2. This parallels the functional differences in conferring PB responsiveness between NR1 and NR3 on the one hand, and NR2 on the other.
Figures



Similar articles
-
Mutational analysis of the CYP2B2 phenobarbital response unit and inhibitory effect of the constitutive androstane receptor on phenobarbital responsiveness.J Biol Chem. 2000 Dec 8;275(49):38427-36. doi: 10.1074/jbc.M005776200. J Biol Chem. 2000. PMID: 10993889
-
Analysis of multiple nuclear receptor binding sites for CAR/RXR in the phenobarbital responsive unit of CYP2B2.Arch Biochem Biophys. 2006 Jul 15;451(2):119-27. doi: 10.1016/j.abb.2006.04.016. Epub 2006 May 8. Arch Biochem Biophys. 2006. PMID: 16725103
-
Chromatin assembly enhances binding to the CYP2B1 phenobarbital-responsive unit (PBRU) of nuclear factor-1, which binds simultaneously with constitutive androstane receptor (CAR)/retinoid X receptor (RXR) and enhances CAR/RXR-mediated activation of the PBRU.J Biol Chem. 2001 Mar 9;276(10):7559-67. doi: 10.1074/jbc.M008090200. Epub 2000 Dec 11. J Biol Chem. 2001. PMID: 11113125
-
Phenobarbital-induced expression of cytochrome P450 genes.Acta Biochim Pol. 2000;47(4):1093-105. Acta Biochim Pol. 2000. PMID: 11996099 Review.
-
Phenobarbital induction of drug/steroid-metabolizing enzymes and nuclear receptor CAR.Biochim Biophys Acta. 2003 Feb 17;1619(3):239-42. doi: 10.1016/s0304-4165(02)00482-8. Biochim Biophys Acta. 2003. PMID: 12573483 Review.
Cited by
-
Thyroid hormone receptor alpha 1-beta 1 expression in epididymal epithelium from euthyroid and hypothyroid rats.Histochem Cell Biol. 2008 May;129(5):631-42. doi: 10.1007/s00418-008-0397-8. Epub 2008 Feb 26. Histochem Cell Biol. 2008. PMID: 18299881
-
A proximity-based graph clustering method for the identification and application of transcription factor clusters.BMC Bioinformatics. 2017 Nov 29;18(1):530. doi: 10.1186/s12859-017-1935-y. BMC Bioinformatics. 2017. PMID: 29187152 Free PMC article.
References
-
- Guengerich F. P. Boca Raton: CRC Press; 1987. Mammalian Cytochromes P-450, vol. 1.
-
- Nebert D. W., Russell D. W. Clinical importance of the cytochromes P450. Lancet. 2002;360:1155–1162. - PubMed
-
- Okey A. B. Enzyme induction in the cytochrome P-450 system. Pharmacol. Ther. 1990;45:241–298. - PubMed
-
- Handschin C., Meyer U. A. Induction of drug metabolism: the role of nuclear receptors. Pharmacol. Rev. 2003;55:649–673. - PubMed
-
- Handschin C., Meyer U. A. A conserved nuclear receptor consensus sequence (DR-4) mediates transcriptional activation of the chicken CYP2H1 gene by phenobarbital in a hepatoma cell line. J. Biol. Chem. 2000;275:13362–13369. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

