Clonal expansion of hepatocytes during chronic woodchuck hepatitis virus infection
- PMID: 15657132
- PMCID: PMC544623
- DOI: 10.1073/pnas.0409332102
Clonal expansion of hepatocytes during chronic woodchuck hepatitis virus infection
Abstract
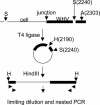
Chronic hepadnavirus infections cause liver damage with ongoing death and regeneration of hepatocytes. In the present study we set out to quantify the extent of liver turnover by measuring the clonal proliferation of hepatocytes by using integrated viral DNA as a genetic marker for individual hepatocyte lineages. Liver tissue from woodchucks with chronic woodchuck hepatitis virus (WHV) infection was assayed for randomly integrated viral DNA by using inverse PCR. Serial endpoint dilution of viral-cell junction fragments into 96-well plates, followed by nested PCR and DNA sequencing, was used to determine the copy number of specific viral cell junctions as a measure of the clonal distribution of infected cell subpopulations. The results indicated that the livers contained a minimum of 100,000 clones of >1,000 cells containing integrated DNA, representing at least 0.2% of the hepatocyte population of the liver. Because cells with integrated WHV DNA comprised only 1-2% of total liver cells, it is likely that the total number of clones far exceeds this estimate, with as much as one-half of the liver derived from high copy clones of >1,000 cells. It may be inferred that these clones have a strong selective growth or survival advantage. The results provide evidence for a large amount of hepatocyte proliferation and selection having occurred during the period of chronic WHV infection ( approximately 1.5 years) in these animals.
Figures



Similar articles
-
Hepatitis B virus cccDNA clearance: killing for curing?Hepatology. 2005 Dec;42(6):1453-5. doi: 10.1002/hep.20976. Hepatology. 2005. PMID: 16317676
-
Replication of naturally occurring woodchuck hepatitis virus deletion mutants in primary hepatocyte cultures and after transmission to naive woodchucks.J Virol. 2001 Apr;75(8):3811-8. doi: 10.1128/JVI.75.8.3811-3818.2001. J Virol. 2001. PMID: 11264370 Free PMC article.
-
The amount of hepatocyte turnover that occurred during resolution of transient hepadnavirus infections was lower when virus replication was inhibited with entecavir.J Virol. 2009 Feb;83(4):1778-89. doi: 10.1128/JVI.01587-08. Epub 2008 Dec 10. J Virol. 2009. PMID: 19073743 Free PMC article.
-
Clonal expansion of hepatocytes with a selective advantage occurs during all stages of chronic hepatitis B virus infection.J Viral Hepat. 2015 Sep;22(9):737-53. doi: 10.1111/jvh.12380. Epub 2015 Jan 26. J Viral Hepat. 2015. PMID: 25619231
-
Hepatitis B Virus DNA Integration and Clonal Expansion of Hepatocytes in the Chronically Infected Liver.Viruses. 2021 Jan 30;13(2):210. doi: 10.3390/v13020210. Viruses. 2021. PMID: 33573130 Free PMC article. Review.
Cited by
-
Persistence of hepatitis B virus covalently closed circular DNA in hepatocytes: molecular mechanisms and clinical significance.Emerg Microbes Infect. 2014 Sep;3(9):e64. doi: 10.1038/emi.2014.64. Epub 2014 Sep 17. Emerg Microbes Infect. 2014. PMID: 26038757 Free PMC article. Review.
-
The liver of woodchucks chronically infected with the woodchuck hepatitis virus contains foci of virus core antigen-negative hepatocytes with both altered and normal morphology.Virology. 2007 Mar 15;359(2):283-94. doi: 10.1016/j.virol.2006.09.034. Epub 2006 Oct 31. Virology. 2007. PMID: 17078989 Free PMC article.
-
The Role of cccDNA in HBV Maintenance.Viruses. 2017 Jun 21;9(6):156. doi: 10.3390/v9060156. Viruses. 2017. PMID: 28635668 Free PMC article. Review.
-
Hepatitis B Viral Protein HBx: Roles in Viral Replication and Hepatocarcinogenesis.Viruses. 2024 Aug 26;16(9):1361. doi: 10.3390/v16091361. Viruses. 2024. PMID: 39339838 Free PMC article. Review.
-
Two fragments of HBV DNA integrated into chrX: 11009033 and its genetic regulation in HepG2.2.15.Mol Med Rep. 2023 May;27(5):98. doi: 10.3892/mmr.2023.12985. Epub 2023 Mar 24. Mol Med Rep. 2023. PMID: 36960866 Free PMC article.
References
-
- Beasley, R. P. (1982) Hepatology 2, 21S–26S.
-
- Vemura, R. P., Aragona, E. & Gupta, S. (1992) Hepatology 16, 968–973. - PubMed
-
- Seki, S., Sakaguchi, H., Kawakita, N., Yanai, A., Kuroki, T., Mizoguchi, Y., Kobayashi, K. & Monna, T. (1991) Hepatology 14, 781–788. - PubMed
-
- MacDonald, R. A. (1960) Arch. Int. Med. 107, 335–343. - PubMed
-
- Grisham, J. W. (1962) Cancer Res. 22, 842–849. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

