Mechanisms of transcription-replication collisions in bacteria
- PMID: 15657418
- PMCID: PMC544003
- DOI: 10.1128/MCB.25.3.888-895.2005
Mechanisms of transcription-replication collisions in bacteria
Abstract
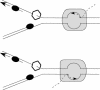
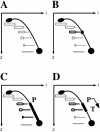
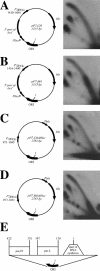
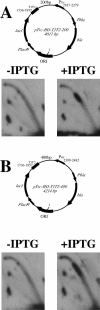
While collisions between replication and transcription in bacteria are deemed inevitable, the fine details of the interplay between the two machineries are poorly understood. In this study, we evaluate the effects of transcription on the replication fork progression in vivo, by using electrophoresis analysis of replication intermediates. Studying Escherichia coli plasmids, which carry constitutive or inducible promoters in different orientations relative to the replication origin, we show that the mutual orientation of the two processes determines their mode of interaction. Replication elongation appears not to be affected by transcription proceeding in the codirectional orientation. Head-on transcription, by contrast, leads to severe inhibition of the replication fork progression. Furthermore, we evaluate the mechanism of this inhibition by limiting the area of direct contact between the two machineries. We observe that replication pausing zones coincide exactly with transcribed DNA segments. We conclude, therefore, that the replication fork is most likely attenuated upon direct physical interaction with the head-on transcription machinery.
Figures

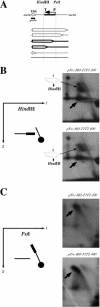
References
-
- Aman, E., B. Ochs, and K.-J. Abel. 1988. Tightly regulated tac promoter vectors useful for the expression of unfused and fused proteins in Escherichia coli. Gene 69:301-315. - PubMed
-
- Blattner, F. R., G. Plunkett III, C. A. Bloch, N. T. Perna, V. Burland, M. Riley, J. Collado-Vides, J. D. Glasner, C. K. Rode, G. F. Mayhew, J. Gregor, N. W. Davis, H. A. Kirkpatrick, M. A. Goeden, D. J. Rose, B. Mau, and Y. Shao. 1997. The complete genome sequence of Escherichia coli K-12. Science 277:1453-1474. - PubMed
-
- Brewer, B. J. 1988. When polymerases collide: replication and the transcriptional organization of the E. coli chromosome. Cell 53:679-686. - PubMed
-
- Brewer, B. J., and W. L. Fangman. 1987. The localization of replication origins on ARS plasmids in S. cerevisiae. Cell 51:463-471. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
