Disparate subcellular localization patterns of Pseudomonas aeruginosa Type IV pilus ATPases involved in twitching motility
- PMID: 15659660
- PMCID: PMC545728
- DOI: 10.1128/JB.187.3.829-839.2005
Disparate subcellular localization patterns of Pseudomonas aeruginosa Type IV pilus ATPases involved in twitching motility
Abstract
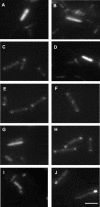
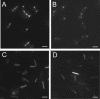
The opportunistic pathogen Pseudomonas aeruginosa expresses polar type IV pili (TFP), which are responsible for adhesion to various materials and twitching motility on surfaces. Twitching occurs by alternate extension and retraction of TFP, which arise from assembly and disassembly of pilin subunits at the base of the pilus. The ATPase PilB promotes pilin assembly, while the ATPase PilT or PilU or both promote pilin dissociation. Fluorescent fusions to two of the three ATPases (PilT and PilU) were functional, as shown by complementation of the corresponding mutants. PilB and PilT fusions localized to both poles, while PilU fusions localized only to the piliated pole. To identify the portion of the ATPases required for localization, sequential C-terminal deletions of PilT and PilU were generated. The conserved His and Walker B boxes were dispensable for polar localization but were required for twitching motility, showing that localization and function could be uncoupled. Truncated fusions that retained polar localization maintained their distinctive distribution patterns. To dissect the cellular factors involved in establishing polarity, fusion protein localization was monitored with a panel of TFP mutants. The localization of yellow fluorescent protein (YFP)-PilT and YFP-PilU was independent of the subunit PilA, other TFP ATPases, and TFP-associated proteins previously shown to be associated with the membrane or exhibiting polar localization. In contrast, YFP-PilB exhibited diffuse cytoplasmic localization in a pilC mutant, suggesting that PilC is required for polar localization of PilB. Finally, localization studies performed with fluorescent ATPase chimeras of PilT and PilU demonstrated that information responsible for the characteristic localization patterns of the ATPases likely resides in their N termini.
Figures




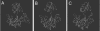

References
-
- Boyd, J. M. 2000. Localization of the histidine kinase PilS to the poles of Pseudomonas aeruginosa and identification of a localization domain. Mol. Microbiol. 36:153-162. - PubMed
-
- Bradley, D. E. 1980. A function of Pseudomonas aeruginosa PAO polar pili: twitching motility. Can. J. Microbiol. 26:146-154. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

