NopB, a type III secreted protein of Rhizobium sp. strain NGR234, is associated with pilus-like surface appendages
- PMID: 15659692
- PMCID: PMC545724
- DOI: 10.1128/JB.187.3.1173-1181.2005
NopB, a type III secreted protein of Rhizobium sp. strain NGR234, is associated with pilus-like surface appendages
Abstract
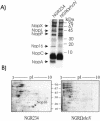
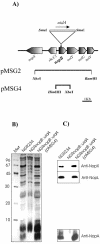
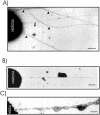
Rhizobium sp. strain NGR234 possesses a functional type three secretion system (TTSS), through which a number of proteins, called nodulation outer proteins (Nops), are delivered to the outside of the cell. A major constraint to the identification of Nops is their low abundance in the supernatants of NGR234 strains grown in culture. To overcome this limitation, a more sensitive proteomics-based strategy was developed. Secreted proteins from wild-type NGR234 were separated by two-dimensional gel electrophoresis, and the gel was compared to similar gels containing the proteins from a TTSS mutant (NGROmegarhcN). To identify the proteins, spots unique to the NGR234 gels were analyzed by matrix-assisted laser desorption ionization-time of flight mass spectrometry and the data were compared to the sequence of the symbiotic plasmid of NGR234. A nonpolar mutant of one of these proteins was generated called NopB. NopB is required for Nop secretion but inhibits the interaction with Pachyrhizus tuberosus and augments nodulation of Tephrosia vogelii. Flavonoids and a functional TTSS are required for the formation of some surface appendages on NGR234. In situ immunogold labeling and isolation of these pili showed that they contain NopB.
Figures




Similar articles
-
NopA is associated with cell surface appendages produced by the type III secretion system of Rhizobium sp. strain NGR234.Mol Plant Microbe Interact. 2005 May;18(5):499-507. doi: 10.1094/MPMI-18-0499. Mol Plant Microbe Interact. 2005. PMID: 15915648
-
Characterization of Nops, nodulation outer proteins, secreted via the type III secretion system of NGR234.Mol Plant Microbe Interact. 2003 Sep;16(9):743-51. doi: 10.1094/MPMI.2003.16.9.743. Mol Plant Microbe Interact. 2003. PMID: 12971597
-
Characterization of NopP, a type III secreted effector of Rhizobium sp. strain NGR234.J Bacteriol. 2004 Jul;186(14):4774-80. doi: 10.1128/JB.186.14.4774-4780.2004. J Bacteriol. 2004. PMID: 15231809 Free PMC article.
-
Nodulation outer proteins: double-edged swords of symbiotic rhizobia.Biochem J. 2015 Sep 15;470(3):263-74. doi: 10.1042/BJ20150518. Biochem J. 2015. PMID: 26341483 Review.
-
Molecular aspects of soybean cultivar-specific nodulation by Sinorhizobium fredii USDA257.Indian J Exp Biol. 2003 Oct;41(10):1114-23. Indian J Exp Biol. 2003. PMID: 15242277 Review.
Cited by
-
Secretion systems and signal exchange between nitrogen-fixing rhizobia and legumes.Front Plant Sci. 2015 Jul 1;6:491. doi: 10.3389/fpls.2015.00491. eCollection 2015. Front Plant Sci. 2015. PMID: 26191069 Free PMC article. Review.
-
Tailoring plant-associated microbial inoculants in agriculture: a roadmap for successful application.J Exp Bot. 2020 Jun 26;71(13):3878-3901. doi: 10.1093/jxb/eraa111. J Exp Bot. 2020. PMID: 32157287 Free PMC article. Review.
-
Commonalities and differences of T3SSs in rhizobia and plant pathogenic bacteria.Front Plant Sci. 2014 Mar 27;5:114. doi: 10.3389/fpls.2014.00114. eCollection 2014. Front Plant Sci. 2014. PMID: 24723933 Free PMC article. Review.
-
Rhizobial measures to evade host defense strategies and endogenous threats to persistent symbiotic nitrogen fixation: a focus on two legume-rhizobium model systems.Cell Mol Life Sci. 2011 Apr;68(8):1327-39. doi: 10.1007/s00018-011-0650-5. Epub 2011 Mar 2. Cell Mol Life Sci. 2011. PMID: 21365276 Free PMC article. Review.
-
Identification of protein secretion systems and novel secreted proteins in Rhizobium leguminosarum bv. viciae.BMC Genomics. 2008 Jan 29;9:55. doi: 10.1186/1471-2164-9-55. BMC Genomics. 2008. PMID: 18230162 Free PMC article.
References
-
- Aizawa, S. I. 2001. Bacterial flagella and type III secretion systems. FEMS Microbiol. Lett. 202:157-164. - PubMed
-
- Ausubel, F., R. Brent, R. Kingston, D. Moore, J. Seidman, and J. Smith. 1991. Current protocols in molecular biology. John Wiley & Sons, New York, N.Y.
-
- Bartsev, A. V., N. M. Boukli, W. J. Deakin, C. Staehelin, and W. Broughton. 2003. Purification and phosphorylation of the effector protein NopL from Rhizobium sp. NGR234. FEBS Lett. 554:271-274. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

