Inducible dissociation of SCF(Met30) ubiquitin ligase mediates a rapid transcriptional response to cadmium
- PMID: 15660125
- PMCID: PMC548661
- DOI: 10.1038/sj.emboj.7600556
Inducible dissociation of SCF(Met30) ubiquitin ligase mediates a rapid transcriptional response to cadmium
Abstract
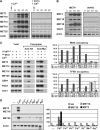
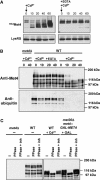
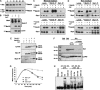
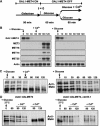
Activity of the Met4 transcription factor is antagonized by the SCF(Met30) ubiquitin ligase by degradation-dependent and degradation-independent mechanisms, in minimal and rich nutrient conditions, respectively. In this study, we show that the heavy metal Cd2+ over-rides both mechanisms to enable rapid Met4-dependent induction of metabolic networks needed for production of the antioxidant and Cd2+-chelating agent glutathione. Cd2+ inhibits SCF(Met30) activity through rapid dissociation of the F-box protein Met30 from the holocomplex. In minimal medium, dissociation of SCF(Met30) complex is sufficient to impair the methionine-induced degradation of Met4. In rich medium, dissociation of the SCF(Met30) complex is accompanied by a deubiquitylation mechanism that rapidly removes inhibitory ubiquitin moieties from Met4. Post-translational control of SCF(Met30) assembly by a physiological stress to allow rapid induction of a protective gene expression program represents a novel mode of regulation in the ubiquitin system.
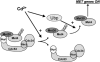
Figures


Similar articles
-
The yeast ubiquitin ligase SCFMet30 regulates heavy metal response.Mol Biol Cell. 2005 Apr;16(4):1872-82. doi: 10.1091/mbc.e04-12-1130. Epub 2005 Feb 2. Mol Biol Cell. 2005. PMID: 15689486 Free PMC article.
-
Determinants of the ubiquitin-mediated degradation of the Met4 transcription factor.J Biol Chem. 2006 Apr 28;281(17):11744-54. doi: 10.1074/jbc.M600037200. Epub 2006 Feb 23. J Biol Chem. 2006. PMID: 16497670
-
A dominant suppressor mutation of the met30 cell cycle defect suggests regulation of the Saccharomyces cerevisiae Met4-Cbf1 transcription complex by Met32.J Biol Chem. 2008 Apr 25;283(17):11615-24. doi: 10.1074/jbc.M708230200. Epub 2008 Feb 28. J Biol Chem. 2008. PMID: 18308733 Free PMC article.
-
Regulation of the cadmium stress response through SCF-like ubiquitin ligases: comparison between Saccharomyces cerevisiae, Schizosaccharomyces pombe and mammalian cells.Biochimie. 2006 Nov;88(11):1673-85. doi: 10.1016/j.biochi.2006.03.001. Epub 2006 Mar 23. Biochimie. 2006. PMID: 16584827 Review.
-
Proteolytic regulation of metabolic enzymes by E3 ubiquitin ligase complexes: lessons from yeast.Crit Rev Biochem Mol Biol. 2015;50(6):489-502. doi: 10.3109/10409238.2015.1081869. Epub 2015 Sep 11. Crit Rev Biochem Mol Biol. 2015. PMID: 26362128 Review.
Cited by
-
Skp1 Independent Function of Cdc53/Cul1 in F-box Protein Homeostasis.PLoS Genet. 2015 Dec 10;11(12):e1005727. doi: 10.1371/journal.pgen.1005727. eCollection 2015 Dec. PLoS Genet. 2015. PMID: 26656496 Free PMC article.
-
Dissection of combinatorial control by the Met4 transcriptional complex.Mol Biol Cell. 2010 Feb 1;21(3):456-69. doi: 10.1091/mbc.e09-05-0420. Epub 2009 Nov 25. Mol Biol Cell. 2010. PMID: 19940020 Free PMC article.
-
Unique role for the UbL-UbA protein Ddi1 in turnover of SCFUfo1 complexes.Mol Cell Biol. 2006 Mar;26(5):1579-88. doi: 10.1128/MCB.26.5.1579-1588.2006. Mol Cell Biol. 2006. PMID: 16478980 Free PMC article.
-
Signal-induced disassembly of the SCF ubiquitin ligase complex by Cdc48/p97.Mol Cell. 2012 Oct 26;48(2):288-97. doi: 10.1016/j.molcel.2012.08.015. Epub 2012 Sep 20. Mol Cell. 2012. PMID: 23000173 Free PMC article.
-
Transcriptional plasticity through differential assembly of a multiprotein activation complex.Nucleic Acids Res. 2010 Aug;38(15):4998-5014. doi: 10.1093/nar/gkq257. Epub 2010 Apr 14. Nucleic Acids Res. 2010. PMID: 20392822 Free PMC article.
References
-
- Blackwell KJ, Tobin JM, Avery SV (1998) Manganese toxicity towards Saccharomyces cerevisiae: dependence on intracellular and extracellular magnesium concentrations. Appl Microbiol Biotechnol 49: 751–757 - PubMed
-
- Cantoni GL (1977) S-adenosylmethionine: present status and future perspectives. In The Biochemistry of S-adenosylmethionine, Salvatore F, Borek E, Zappia V, Williams-Ashman HG, Schlenk F (eds) pp 557–577. New York: Columbia University Press
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

