DNA analysis with multiplex microarray-enhanced PCR
- PMID: 15661850
- PMCID: PMC548369
- DOI: 10.1093/nar/gnh184
DNA analysis with multiplex microarray-enhanced PCR
Abstract
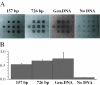
We have developed a highly sensitive method for DNA analysis on 3D gel element microarrays, a technique we call multiplex microarray-enhanced PCR (MME-PCR). Two amplification strategies are carried out simultaneously in the reaction chamber: on or within gel elements, and in bulk solution over the gel element array. MME-PCR is initiated by multiple complex primers containing gene-specific, forward and reverse, sequences appended to the 3' end of a universal amplification primer. The complex primer pair is covalently tethered through its 5' end to the polyacryl- amide backbone. In the bulk solution above the gel element array, a single pair of unattached universal primers simultaneously directs pseudo-monoplex PCR of all targets according to normal solution-phase PCR. The presence of a single universal PCR primer pair in solution accelerates amplification within gel elements and eliminates the problem of primer interference that is common to conventional multiplex PCR. We show 10(6)-fold amplification of targeted DNA after 50 cycles with average amplification efficiency 1.34 per cycle, and demonstrate specific on-chip amplification of six genes in Bacillus subtilis. All six genes were detected at 4.5 pg of bacterial genomic DNA (equivalent to 10(3) genomes) in 60 independent amplification reactions performed simultaneously in single reaction chamber.
Figures

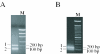
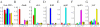
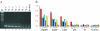
Similar articles
-
A universal multiplex PCR strategy for 100-plex amplification using a hydrophobically patterned microarray.Lab Chip. 2011 Nov 7;11(21):3609-18. doi: 10.1039/c1lc20526a. Epub 2011 Sep 9. Lab Chip. 2011. PMID: 21909519
-
The construction and use of bacterial DNA microarrays based on an optimized two-stage PCR strategy.BMC Genomics. 2003 Jun 12;4(1):23. doi: 10.1186/1471-2164-4-23. Epub 2003 Jun 12. BMC Genomics. 2003. PMID: 12803655 Free PMC article.
-
Multiplex amplification enabled by selective circularization of large sets of genomic DNA fragments.Nucleic Acids Res. 2005 Apr 28;33(8):e71. doi: 10.1093/nar/gni070. Nucleic Acids Res. 2005. PMID: 15860768 Free PMC article.
-
The real-time polymerase chain reaction.Mol Aspects Med. 2006 Apr-Jun;27(2-3):95-125. doi: 10.1016/j.mam.2005.12.007. Epub 2006 Feb 3. Mol Aspects Med. 2006. PMID: 16460794 Review.
-
MultiPrimer: a system for microarray PCR primer design.Methods Mol Biol. 2007;402:305-14. doi: 10.1007/978-1-59745-528-2_15. Methods Mol Biol. 2007. PMID: 17951802 Review.
Cited by
-
Multiple primer extension by DNA polymerase on a novel plastic DNA array coated with a biocompatible polymer.Nucleic Acids Res. 2007;35(1):e3. doi: 10.1093/nar/gkl939. Epub 2006 Nov 28. Nucleic Acids Res. 2007. PMID: 17135189 Free PMC article.
-
Large scale multiplex PCR improves pathogen detection by DNA microarrays.BMC Microbiol. 2009 Jan 3;9:1. doi: 10.1186/1471-2180-9-1. BMC Microbiol. 2009. PMID: 19121223 Free PMC article.
-
Genotyping of KRAS Mutational Status by the In-Check Lab-on-Chip Platform.Sensors (Basel). 2018 Jan 5;18(1):131. doi: 10.3390/s18010131. Sensors (Basel). 2018. PMID: 29304017 Free PMC article.
-
Highly parallel and short-acting amplification with locus-specific primers to detect single nucleotide polymorphisms by the DigiTag2 assay.PLoS One. 2012;7(1):e29967. doi: 10.1371/journal.pone.0029967. Epub 2012 Jan 13. PLoS One. 2012. PMID: 22253840 Free PMC article.
-
The role of DNA diffusion in solid phase polymerase chain reaction with gel-immobilized primers in planar and capillary microarray format.Biomicrofluidics. 2009 Dec 1;3(4):44112. doi: 10.1063/1.3271461. Biomicrofluidics. 2009. PMID: 20216974 Free PMC article.
References
-
- Lipshutz R.J., Fodor S.P., Gingeras T.R., Lockhart D.J. High density synthetic oligonucleotide arrays. Nature Genet. 1999;21((Suppl.)):20–24. - PubMed
-
- Brown P.O., Botstein D. Exploring the new world of the genome with DNA microarrays. Nature Genet. 1999;21((Suppl.)):33–37. - PubMed
-
- Duggan D.J., Bittner M., Chen Y., Meltzer P., Trent J.M. Expression profiling using cDNA microarrays. Nature Genet. 1999;21((Suppl.)):10–14. - PubMed
-
- Edwards M.C., Gibbs R.A. Multiplex PCR: advantages, development, and applications. PCR Methods Appl. 1994;3:S65–S75. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

