CASCAD: a database of annotated candidate single nucleotide polymorphisms associated with expressed sequences
- PMID: 15676075
- PMCID: PMC548278
- DOI: 10.1186/1471-2164-6-10
CASCAD: a database of annotated candidate single nucleotide polymorphisms associated with expressed sequences
Abstract
Background: With the recent progress made in large-scale genome sequencing projects a vast amount of novel data is becoming available. A comparative sequence analysis, exploiting sequence information from various resources, can be used to uncover hidden information, such as genetic variation. Although there are enormous amounts of SNPs for a wide variety of organisms submitted to NCBI dbSNP and annotated in most genome assembly viewers like Ensembl and the UCSC Genome Browser, these platforms do not easily allow for extensive annotation and incorporation of experimental data supporting the polymorphism. However, such information is very important for selecting the most promising and useful candidate polymorphisms for use in experimental setups.
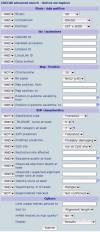
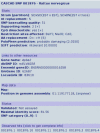
Description: The CASCAD database is designed for presentation and query of candidate SNPs that are retrieved by in silico mining of high-throughput sequencing data. Currently, the database provides collections of laboratory rat (Rattus norvegicus) and zebrafish (Danio rerio) candidate SNPs. The database stores detailed information about raw data supporting the candidate, extensive annotation and links to external databases (e.g. GenBank, Ensembl, UniGene, and LocusLink), verification information, and predictions of a potential effect for non-synonymous polymorphisms in coding regions. The CASCAD website allows search based on an arbitrary combination of 27 different parameters related to characteristics like candidate SNP quality, genomic localization, and sequence data source or strain. In addition, the database can be queried with any custom nucleotide sequences of interest. The interface is crosslinked to other public databases and tightly coupled with primer design and local genome assembly interfaces in order to facilitate experimental verification of candidates.
Conclusions: The CASCAD database discloses detailed information on rat and zebrafish candidate SNPs, including the raw data underlying its discovery. An advanced web-based search interface http://cascad.niob.knaw.nl allows universal access to the database content and allows various queries supporting many types of research utilizing single nucleotide polymorphisms.
Figures
Similar articles
-
SNP mining porcine ESTs with MAVIANT, a novel tool for SNP evaluation and annotation.Bioinformatics. 2007 Jul 1;23(13):i387-91. doi: 10.1093/bioinformatics/btm192. Bioinformatics. 2007. PMID: 17646321
-
Piloting the zebrafish genome browser.Dev Dyn. 2006 Mar;235(3):747-53. doi: 10.1002/dvdy.20661. Dev Dyn. 2006. PMID: 16372332
-
DigiPINS: a database for vertebrate exonic single nucleotide polymorphisms and its application to cancer association studies.Biochimie. 2008 Apr;90(4):563-9. doi: 10.1016/j.biochi.2007.09.017. Epub 2007 Sep 29. Biochimie. 2008. PMID: 17988782
-
Genome information resources - developments at Ensembl.Trends Genet. 2004 Jun;20(6):268-72. doi: 10.1016/j.tig.2004.04.002. Trends Genet. 2004. PMID: 15145580 Review.
-
Sequence assembly.Comput Biol Chem. 2009 Apr;33(2):121-36. doi: 10.1016/j.compbiolchem.2008.11.003. Epub 2008 Dec 6. Comput Biol Chem. 2009. PMID: 19152793 Review.
Cited by
-
A major zebrafish polymorphism resource for genetic mapping.Genome Biol. 2007;8(4):R55. doi: 10.1186/gb-2007-8-4-r55. Genome Biol. 2007. PMID: 17428331 Free PMC article.
-
Genetic variation in the zebrafish.Genome Res. 2006 Apr;16(4):491-7. doi: 10.1101/gr.4791006. Epub 2006 Mar 13. Genome Res. 2006. PMID: 16533913 Free PMC article.
-
Identifying functional single nucleotide polymorphisms in the human CArGome.Physiol Genomics. 2011 Sep 22;43(18):1038-48. doi: 10.1152/physiolgenomics.00098.2011. Epub 2011 Jul 19. Physiol Genomics. 2011. PMID: 21771879 Free PMC article.
-
Seq4SNPs: new software for retrieval of multiple, accurately annotated DNA sequences, ready formatted for SNP assay design.BMC Bioinformatics. 2009 Jun 12;10:180. doi: 10.1186/1471-2105-10-180. BMC Bioinformatics. 2009. PMID: 19523221 Free PMC article.
-
SNP discovery by mismatch-targeting of Mu transposition.Nucleic Acids Res. 2007;35(6):e44. doi: 10.1093/nar/gkm070. Epub 2007 Feb 20. Nucleic Acids Res. 2007. PMID: 17311815 Free PMC article.
References
-
- Primer design interface http://primers.niob.knaw.nl
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources