Structure of the kainate receptor subunit GluR6 agonist-binding domain complexed with domoic acid
- PMID: 15677325
- PMCID: PMC547884
- DOI: 10.1073/pnas.0409573102
Structure of the kainate receptor subunit GluR6 agonist-binding domain complexed with domoic acid
Abstract
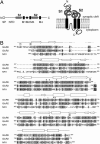
We report the crystal structure of the glycosylated ligand-binding (S1S2) domain of the kainate receptor subunit GluR6, in complex with the agonist domoate. The structure shows the expected overall homology with AMPA and NMDA receptor subunit structures but reveals an unexpected binding mode for the side chain of domoate, in which contact is made to the larger lobe only (lobe I). In common with the AMPA receptor subunit GluR2, the GluR6 S1S2 domain associates as a dimer, with many of the interdimer contacts being conserved. Subtle differences in these contacts provide a structural explanation for why GluR2 L483Y and GluR3 L507Y are nondesensitizing, but GluR6, which has a tyrosine at that site, is not. The structure incorporates native glycosylation, which has not previously been described for ionotropic glutamate receptors. The position of the sugars near the subunit interface rules out their direct involvement in subunit association but leaves open the possibility of indirect modulation. Finally, we observed several tetrameric assemblies that satisfy topological constraints with respect to connection to the receptor pore, and which are therefore candidates for the native quaternary structure.
Figures



Similar articles
-
Interface interactions modulating desensitization of the kainate-selective ionotropic glutamate receptor subunit GluR6.J Neurosci. 2006 Sep 27;26(39):10033-42. doi: 10.1523/JNEUROSCI.2750-06.2006. J Neurosci. 2006. PMID: 17005866 Free PMC article.
-
Crystal structure of the kainate receptor GluR5 ligand-binding core in complex with (S)-glutamate.FEBS Lett. 2005 Feb 14;579(5):1154-60. doi: 10.1016/j.febslet.2005.01.012. FEBS Lett. 2005. PMID: 15710405
-
Crystal structures of the GluR5 and GluR6 ligand binding cores: molecular mechanisms underlying kainate receptor selectivity.Neuron. 2005 Feb 17;45(4):539-52. doi: 10.1016/j.neuron.2005.01.031. Neuron. 2005. PMID: 15721240
-
Lessons from crystal structures of kainate receptors.Neuropharmacology. 2017 Jan;112(Pt A):16-28. doi: 10.1016/j.neuropharm.2016.05.014. Epub 2016 May 26. Neuropharmacology. 2017. PMID: 27236079 Review.
-
Stereostructure-activity studies on agonists at the AMPA and kainate subtypes of ionotropic glutamate receptors.Chirality. 2003 Feb;15(2):167-79. doi: 10.1002/chir.10177. Chirality. 2003. PMID: 12520509 Review.
Cited by
-
Correlating efficacy and desensitization with GluK2 ligand-binding domain movements.Open Biol. 2013 May 29;3(5):130051. doi: 10.1098/rsob.130051. Open Biol. 2013. PMID: 23720540 Free PMC article.
-
Cations but not anions regulate the responsiveness of kainate receptors.J Neurosci. 2011 Feb 9;31(6):2136-44. doi: 10.1523/JNEUROSCI.4314-10.2011. J Neurosci. 2011. PMID: 21307250 Free PMC article.
-
Structure of the S1S2 glutamate binding domain of GLuR3.Proteins. 2009 May 15;75(3):628-37. doi: 10.1002/prot.22274. Proteins. 2009. PMID: 19003990 Free PMC article.
-
Structural and Functional Insights into GluK3-kainate Receptor Desensitization and Recovery.Sci Rep. 2019 Jul 16;9(1):10254. doi: 10.1038/s41598-019-46770-z. Sci Rep. 2019. PMID: 31311973 Free PMC article.
-
Binding site and ligand flexibility revealed by high resolution crystal structures of GluK1 competitive antagonists.Neuropharmacology. 2011 Jan;60(1):126-34. doi: 10.1016/j.neuropharm.2010.06.002. Epub 2010 Jun 15. Neuropharmacology. 2011. PMID: 20558186 Free PMC article.
References
-
- Dingledine, R., Borges, K., Bowie, D. & Traynelis, S. F. (1999) Pharmacol. Rev. 51, 7–61. - PubMed
-
- Huettner, J. E. (2003) Prog. Neurobiol. 70, 387–407. - PubMed
-
- Stern-Bach, Y., Bettler, B., Hartley, M., Sheppard, P. O., O'Hara, P. J. & Heinemann, S. F. (1994) Neuron 13, 1345–1357. - PubMed
-
- Armstrong, N., Sun, Y., Chen, G.-Q. & Gouaux, E. (1998) Nature 395, 913–917. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

