A non-ATP-competitive inhibitor of BCR-ABL overrides imatinib resistance
- PMID: 15677719
- PMCID: PMC546016
- DOI: 10.1073/pnas.0408283102
A non-ATP-competitive inhibitor of BCR-ABL overrides imatinib resistance
Erratum in
- Proc Natl Acad Sci U S A. 2005 Apr 12;102(15):5635
Abstract
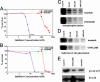
Imatinib, which is an inhibitor of the BCR-ABL tyrosine kinase, has been a remarkable success for the treatment of Philadelphia chromosome-positive (Ph+) chronic myelogenous leukemias (CMLs). However, a significant proportion of patients chronically treated with imatinib develop resistance because of the acquisition of mutations in the kinase domain of BCR-ABL. Mutations occur at residues directly implicated in imatinib binding or, more commonly, at residues important for the ability of the kinase to adopt the specific closed (inactive) conformation to which imatinib binds. In our quest to develop new BCR-ABL inhibitors, we chose to target regions outside the ATP-binding site of this enzyme because these compounds offer the potential to be unaffected by mutations that make CML cells resistant to imatinib. Here we describe the activity of one compound, ON012380, that can specifically inhibit BCR-ABL and induce cell death of Ph+ CML cells at a concentration of <10 nM. Kinetic studies demonstrate that this compound is not ATP-competitive but is substrate-competitive and works synergistically with imatinib in wild-type BCR-ABL inhibition. More importantly, ON012380 was found to induce apoptosis of all of the known imatinib-resistant mutants at concentrations of <10 nM concentration in vitro and cause regression of leukemias induced by i.v. injection of 32Dcl3 cells expressing the imatinib-resistant BCR-ABL isoform T315I. Daily i.v. dosing for up to 3 weeks with a >100 mg/kg concentration of this agent is well tolerated in rodents, without any hematotoxicity.
Figures
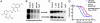




Similar articles
-
Changes associated with the development of resistance to imatinib (STI571) in two leukemia cell lines expressing p210 Bcr/Abl protein.Cancer. 2004 Apr 1;100(7):1459-71. doi: 10.1002/cncr.20131. Cancer. 2004. PMID: 15042680
-
Novel targeted therapies to overcome imatinib mesylate resistance in chronic myeloid leukemia (CML).Crit Rev Oncol Hematol. 2006 Feb;57(2):145-64. doi: 10.1016/j.critrevonc.2005.06.007. Epub 2005 Oct 5. Crit Rev Oncol Hematol. 2006. PMID: 16213151 Review.
-
Sorafenib induces apoptosis specifically in cells expressing BCR/ABL by inhibiting its kinase activity to activate the intrinsic mitochondrial pathway.Cancer Res. 2009 May 1;69(9):3927-36. doi: 10.1158/0008-5472.CAN-08-2978. Epub 2009 Apr 14. Cancer Res. 2009. PMID: 19366808
-
Resistance of Philadelphia-chromosome positive leukemia towards the kinase inhibitor imatinib (STI571, Glivec): a targeted oncoprotein strikes back.Leukemia. 2003 May;17(5):829-38. doi: 10.1038/sj.leu.2402889. Leukemia. 2003. PMID: 12750693 Review.
-
Molecular and chromosomal mechanisms of resistance to imatinib (STI571) therapy.Leukemia. 2002 Nov;16(11):2190-6. doi: 10.1038/sj.leu.2402741. Leukemia. 2002. PMID: 12399961 Clinical Trial.
Cited by
-
The second generation of BCR-ABL tyrosine kinase inhibitors.Int J Hematol. 2006 May;83(4):294-300. doi: 10.1532/IJH97.06025. Int J Hematol. 2006. PMID: 16757427 Review.
-
Imatinib-resistant chronic myeloid leukemia (CML): Current concepts on pathogenesis and new emerging pharmacologic approaches.Biologics. 2007 Dec;1(4):433-48. Biologics. 2007. PMID: 19707313 Free PMC article.
-
Exploring the chemotherapeutic potential of currently used kinase inhibitors: An update.Front Pharmacol. 2023 Jan 9;13:1064472. doi: 10.3389/fphar.2022.1064472. eCollection 2022. Front Pharmacol. 2023. PMID: 36699049 Free PMC article. Review.
-
Molecular biology of the novel anticancer medications: a focus on kinases inhibitors, biologics and CAR T-cell therapy.Inflamm Res. 2025 Feb 17;74(1):41. doi: 10.1007/s00011-025-02008-5. Inflamm Res. 2025. PMID: 39960501 Review.
-
Novel pyrrolo-1,5-benzoxazepine compounds display significant activity against resistant chronic myeloid leukaemia cells in vitro, in ex vivo patient samples and in vivo.Br J Cancer. 2010 May 11;102(10):1474-82. doi: 10.1038/sj.bjc.6605670. Epub 2010 Apr 20. Br J Cancer. 2010. PMID: 20407438 Free PMC article.
References
-
- Nowell, P. C. & Hungerford, D. A. (1960) Science 132, 1497.
-
- Rowley, J. D. (1973) Nature 243, 290–293. - PubMed
-
- DeKlein, A., Van Kessel, A. G., Grosveld, G., Bartram, C. R., Hagemeijer, A., Bostooma, D., Spurr, N. K., Heisterkamp, N., Groffen, J. & Stephenson, J. R. (1982) Nature 300, 765–767. - PubMed
-
- Heisterkamp, N., Stam, K., Groffen, J., de Klein, A. & Grosveld, G. (1985) Nature 315, 758–761. - PubMed
-
- Druker, B. J., Tamura, S., Buchdunger, E., Ohno, S., Segal, G. M., Fanning, S., Zimmermann, J. & Lydon, N. B. (1996) Nat. Med. 2, 561–566. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

