Structural characterization of the highly conserved 98-base sequence at the 3' end of HCV RNA genome and the complementary sequence located at the 5' end of the replicative viral strand
- PMID: 15681619
- PMCID: PMC548360
- DOI: 10.1093/nar/gki218
Structural characterization of the highly conserved 98-base sequence at the 3' end of HCV RNA genome and the complementary sequence located at the 5' end of the replicative viral strand
Abstract
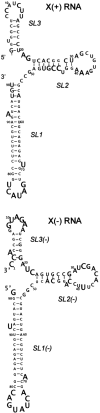
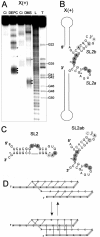
Oligoribonucleotides that corresponded to the X regions of the (+) and (-) polarity strands of HCV RNA, as well as several shorter oligomers comprising defined stem-loop motifs of their predicted secondary structure models, were analyzed by Pb2+-induced cleavage, partial digestion with specific nucleases and chemical modification. Patterns characteristic of the motifs were compared with those obtained for the full-length molecules and on the basis of such 'structural fingerprinting' conclusions concerning folding of regions X were formulated. It turned out that the secondary structure model of X(+) RNA proposed earlier, the three-stem-loop model composed of hairpins SL1, SL2 and SL3, was only partially consistent with our experimental data. We confirmed the presence of SL1 and SL3 motifs and showed that the single-stranded stretch adjacent to the earlier proposed hairpin SL2 contributed to the folding of that region. It seemed to be arranged into two hairpins, which might form a hypothetical pseudoknot by changing their base-pairing systems. These data were discussed in terms of their possible biological significance. On the other hand, analysis of the X(-) RNA and its sub-fragments supported a three-stem-loop secondary structure model for this RNA.
Figures


Similar articles
-
Heterogeneous nuclear ribonucleoprotein I (hnRNP-I/PTB) selectively binds the conserved 3' terminus of hepatitis C viral RNA.Biochem Biophys Res Commun. 1999 Jan 19;254(2):351-62. doi: 10.1006/bbrc.1998.9949. Biochem Biophys Res Commun. 1999. PMID: 9918842
-
Structural domains of the 3'-terminal sequence of the hepatitis C virus replicative strand.Biochemistry. 2008 Nov 18;47(46):12197-207. doi: 10.1021/bi800348g. Epub 2008 Oct 24. Biochemistry. 2008. PMID: 18947238
-
NMR structure of stem-loop SL2 of the HIV-1 psi RNA packaging signal reveals a novel A-U-A base-triple platform.J Mol Biol. 2000 May 26;299(1):145-56. doi: 10.1006/jmbi.2000.3710. J Mol Biol. 2000. PMID: 10860728
-
[Structure and function of the non-coding regions of hepatitis C viral RNA].Postepy Biochem. 2006;52(1):62-71. Postepy Biochem. 2006. PMID: 16869303 Review. Polish.
-
[Highly conserved secondary structures predicted in 5' noncoding region of HCV: a possible connection to viral replication].Nihon Rinsho. 1995 Sep;53 Suppl(Pt 1):43-50. Nihon Rinsho. 1995. PMID: 7563781 Review. Japanese. No abstract available.
Cited by
-
Nuclear Magnetic Resonance Study of RNA Structures at the 3'-End of the Hepatitis C Virus Genome.Biochemistry. 2017 Sep 19;56(37):4972-4984. doi: 10.1021/acs.biochem.7b00573. Epub 2017 Sep 6. Biochemistry. 2017. PMID: 28829576 Free PMC article.
-
New RNA Structural Elements Identified in the Coding Region of the Coxsackie B3 Virus Genome.Viruses. 2020 Oct 30;12(11):1232. doi: 10.3390/v12111232. Viruses. 2020. PMID: 33143071 Free PMC article.
-
The conserved 3'X terminal domain of hepatitis C virus genomic RNA forms a two-stem structure that promotes viral RNA dimerization.Nucleic Acids Res. 2015 Sep 30;43(17):8529-39. doi: 10.1093/nar/gkv786. Epub 2015 Aug 3. Nucleic Acids Res. 2015. PMID: 26240378 Free PMC article.
-
Three-dimensional structure of the 3'X-tail of hepatitis C virus RNA in monomeric and dimeric states.RNA. 2017 Sep;23(9):1465-1476. doi: 10.1261/rna.060632.117. Epub 2017 Jun 19. RNA. 2017. PMID: 28630140 Free PMC article.
-
The low-resolution structural models of hepatitis C virus RNA subdomain 5BSL3.2 and its distal complex with domain 3'X point to conserved regulatory mechanisms within the Flaviviridae family.Nucleic Acids Res. 2022 Feb 28;50(4):2287-2301. doi: 10.1093/nar/gkac061. Nucleic Acids Res. 2022. PMID: 35137150 Free PMC article.
References
-
- World Health Organization. Hepatitis C: global prevalence (update) Weakly Epidemiol. Rec. 1999;49:425–427. - PubMed
-
- Reed K.E., Rice C.M. Overview of hepatitis C virus genome structure, polyprotein processing, and protein properties. Curr. Top. Microbiol. Immunol. 2000;242:55–84. - PubMed
-
- Branch A.D. Hepatitis C virus RNA codes for proteins and replicates: does it also trigger the interferon response? Semin. Liver Dis. 2000;20:57–68. - PubMed
-
- Penin F., Dubuisson J., Rey F.A., Moradpour D., Pawlotsky J.M. Structural biology of hepatitis C virus. Hepatology. 2004;39:5–19. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

