Does water play a structural role in the folding of small nucleic acids?
- PMID: 15681648
- PMCID: PMC1305348
- DOI: 10.1529/biophysj.104.055087
Does water play a structural role in the folding of small nucleic acids?
Abstract
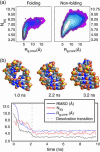
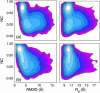
Nucleic acid structure and dynamics are known to be closely coupled to local environmental conditions and, in particular, to the ionic character of the solvent. Here we consider what role the discrete properties of water and ions play in the collapse and folding of small nucleic acids. We study the folding of an experimentally well-characterized RNA hairpin-loop motif (sequence 5'-GGGC[GCAA]GCCU-3') via ensemble molecular dynamics simulation and, with nearly 500 micros of aggregate simulation time using an explicit representation of the ionic solvent, report successful ensemble folding simulations with a predicted folding time of 8.8(+/-2.0) micros, in agreement with experimental measurements of approximately 10 micros. Comparing our results to previous folding simulations using the GB/SA continuum solvent model shows that accounting for water-mediated interactions is necessary to accurately characterize the free energy surface and stochastic nature of folding. The formation of the secondary structure appears to be more rapid than the fastest ionic degrees of freedom, and counterions do not participate discretely in observed folding events. We find that hydrophobic collapse follows a predominantly expulsive mechanism in which a diffusion-search of early structural compaction is followed by the final formation of native structure that occurs in tandem with solvent evacuation.
Figures



Similar articles
-
RNA simulations: probing hairpin unfolding and the dynamics of a GNRA tetraloop.J Mol Biol. 2002 Apr 5;317(4):493-506. doi: 10.1006/jmbi.2002.5447. J Mol Biol. 2002. PMID: 11955005
-
Insights into nucleic acid conformational dynamics from massively parallel stochastic simulations.Biophys J. 2003 Aug;85(2):790-803. doi: 10.1016/S0006-3495(03)74520-2. Biophys J. 2003. PMID: 12885628 Free PMC article.
-
Beta-hairpin folding simulations in atomistic detail using an implicit solvent model.J Mol Biol. 2001 Oct 12;313(1):151-69. doi: 10.1006/jmbi.2001.5033. J Mol Biol. 2001. PMID: 11601853
-
Molecular modeling of nucleic Acid structure: electrostatics and solvation.Curr Protoc Nucleic Acid Chem. 2014 Dec 19;55:7.9.1-27. doi: 10.1002/0471142700.nc0709s55. Curr Protoc Nucleic Acid Chem. 2014. PMID: 25631536 Review.
-
Protein and peptide folding explored with molecular simulations.Acc Chem Res. 2002 Jun;35(6):447-54. doi: 10.1021/ar0100172. Acc Chem Res. 2002. PMID: 12069630 Review.
Cited by
-
Constructing multi-resolution Markov State Models (MSMs) to elucidate RNA hairpin folding mechanisms.Pac Symp Biocomput. 2010:228-39. doi: 10.1142/9789814295291_0025. Pac Symp Biocomput. 2010. PMID: 19908375 Free PMC article.
-
High-resolution reversible folding of hyperstable RNA tetraloops using molecular dynamics simulations.Proc Natl Acad Sci U S A. 2013 Oct 15;110(42):16820-5. doi: 10.1073/pnas.1309392110. Epub 2013 Sep 16. Proc Natl Acad Sci U S A. 2013. PMID: 24043821 Free PMC article. Review.
-
Thermodynamics and kinetics of the hairpin ribozyme from atomistic folding/unfolding simulations.J Mol Biol. 2011 Sep 2;411(5):1128-44. doi: 10.1016/j.jmb.2011.06.042. Epub 2011 Jun 30. J Mol Biol. 2011. PMID: 21740912 Free PMC article.
-
Equilibrium conformational dynamics in an RNA tetraloop from massively parallel molecular dynamics.Nucleic Acids Res. 2010 Aug;38(14):4856-67. doi: 10.1093/nar/gkq134. Epub 2010 Mar 11. Nucleic Acids Res. 2010. PMID: 20223768 Free PMC article.
-
Rapid equilibrium sampling initiated from nonequilibrium data.Proc Natl Acad Sci U S A. 2009 Nov 24;106(47):19765-9. doi: 10.1073/pnas.0909088106. Epub 2009 Sep 29. Proc Natl Acad Sci U S A. 2009. PMID: 19805023 Free PMC article.
References
-
- Berendsen, H. J. C., J. P. M. Postma, W. F. Vangunsteren, A. Dinola, and J. R. Haak. 1984. Molecular-dynamics with coupling to an external bath. J. Chem. Phys. 81:3684–3690.
-
- Cornell, W. D., P. Cieplak, C. I. Bayly, I. R. Gould, K. M. Merz, D. M. Ferguson, D. C. Spellmeyer, T. Fox, J. W. Caldwell, and P. A. Kollman. 1995. A second generation force field for the simulation of proteins, nucleic acids, and organic molecules. J. Am. Chem. Soc. 117:5179–5197.
-
- Darden, T., D. York, and L. Pedersen. 1995. A smooth particle mesh Ewald potential. J. Chem. Phys. 103:3014–3021.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

