Ace2p controls the expression of genes required for cell separation in Schizosaccharomyces pombe
- PMID: 15689498
- PMCID: PMC1073678
- DOI: 10.1091/mbc.e04-06-0442
Ace2p controls the expression of genes required for cell separation in Schizosaccharomyces pombe
Abstract
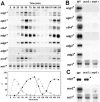
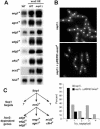
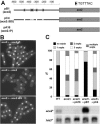
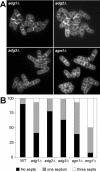
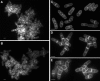
Schizosaccharomyces pombe cells divide by medial fission through contraction of an actomyosin ring and deposition of a multilayered division septum that must be cleaved to release the two daughter cells. Here we describe the identification of seven genes (adg1(+), adg2(+), adg3(+), cfh4(+), agn1(+), eng1(+), and mid2(+)) whose expression is induced by the transcription factor Ace2p. The expression of all of these genes varied during the cell cycle, maximum transcription being observed during septation. At least three of these proteins (Eng1p, Agn1p, and Cfh4p) localize to a ring-like structure that surrounds the septum region during cell separation. Deletion of the previously uncharacterized genes was not lethal to the cells, but produced defects or delays in cell separation to different extents. Electron microscopic observation of mutant cells indicated that the most severe defect is found in eng1Delta agn1Delta cells, lacking the Eng1p endo-beta-1,3-glucanase and the Agn1p endo-alpha-glucanase. The phenotype of this mutant closely resembled that of ace2Delta mutants, forming branched chains of cells. This suggests that these two proteins are the main activities required for cell separation to be completed.
Figures





Similar articles
-
The endo-beta-1,3-glucanase eng1p is required for dissolution of the primary septum during cell separation in Schizosaccharomyces pombe.J Cell Sci. 2003 May 1;116(Pt 9):1689-98. doi: 10.1242/jcs.00377. J Cell Sci. 2003. PMID: 12665550
-
Role of the alpha-glucanase Agn1p in fission-yeast cell separation.Mol Biol Cell. 2004 Aug;15(8):3903-14. doi: 10.1091/mbc.e04-04-0319. Epub 2004 Jun 11. Mol Biol Cell. 2004. PMID: 15194814 Free PMC article.
-
The alpha-glucanase Agn1p is required for cell separation in Schizosaccharomyces pombe.Biol Cell. 2005 Jul;97(7):569-76. doi: 10.1042/BC20040096. Biol Cell. 2005. PMID: 15850449
-
Splitting of the fission yeast septum.FEMS Yeast Res. 2007 Sep;7(6):761-70. doi: 10.1111/j.1567-1364.2007.00266.x. Epub 2007 Jun 27. FEMS Yeast Res. 2007. PMID: 17596184 Review.
-
A transcriptional pathway for cell separation in fission yeast.Cell Cycle. 2005 Jan;4(1):39-41. doi: 10.4161/cc.4.1.1336. Epub 2005 Jan 29. Cell Cycle. 2005. PMID: 15611619 Review.
Cited by
-
Cdk1 activity acts as a quantitative platform for coordinating cell cycle progression with periodic transcription.Nat Commun. 2016 Apr 5;7:11161. doi: 10.1038/ncomms11161. Nat Commun. 2016. PMID: 27045731 Free PMC article.
-
Chitinases Are Essential for Cell Separation in Ustilago maydis.Eukaryot Cell. 2015 Sep;14(9):846-57. doi: 10.1128/EC.00022-15. Epub 2015 May 1. Eukaryot Cell. 2015. PMID: 25934689 Free PMC article.
-
Comprehensive proteomics analysis reveals new substrates and regulators of the fission yeast clp1/cdc14 phosphatase.Mol Cell Proteomics. 2013 May;12(5):1074-86. doi: 10.1074/mcp.M112.025924. Epub 2013 Jan 7. Mol Cell Proteomics. 2013. PMID: 23297348 Free PMC article.
-
Functional characterization of fission yeast transcription factors by overexpression analysis.Genetics. 2013 Aug;194(4):873-84. doi: 10.1534/genetics.113.150870. Epub 2013 May 20. Genetics. 2013. PMID: 23695302 Free PMC article.
-
Adaptation to Industrial Stressors Through Genomic and Transcriptional Plasticity in a Bioethanol Producing Fission Yeast Isolate.G3 (Bethesda). 2020 Apr 9;10(4):1375-1391. doi: 10.1534/g3.119.400986. G3 (Bethesda). 2020. PMID: 32086247 Free PMC article.
References
-
- Bähler, J., Wu, J. Q., Longtine, M. S., Shah, N. G., McKenzie, A., Steever, A. B., Wach, A., Philippsen, P., and Pringle, J. R. (1998b). Heterologous modules for efficient and versatile PCR-based gene targeting in Schizosaccharomyces pombe. Yeast 14, 943-951. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

