Structural probing of a pathogenic tRNA dimer
- PMID: 15701731
- PMCID: PMC1370715
- DOI: 10.1261/rna.7143305
Structural probing of a pathogenic tRNA dimer
Abstract
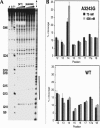
The A3243G mutation within the human mitochondrial (hs mt) tRNALeuUUR gene is associated with maternally inherited deafness and diabetes (MIDD) and other mitochondrial encephalopathies. One of the most pronounced structural effects of this mutation is the disruption of the native structure through stabilization of a high-affinity dimeric complex. We conducted a series of studies that address the structural properties of this tRNA dimer, and we assessed its formation under physiological conditions. Enzymatic probing was used to directly define the dimeric interface for the complex, and a discrete region of the D-stem and loop of hs mt tRNALeuUUR was identified. The dependence of dimerization on magnesium ions and temperature was also tested. The formation of the tRNA dimer is influenced by temperature, with dimerization becoming more efficient at physiological temperature. Complexation of the mutant tRNA is also affected by the amount of magnesium present, and occurs at concentrations present intracellularly. Terbium probing experiments revealed a specific metal ion-binding site localized at the site of the A3243G mutation that is unique to the dimer structure. This metal ion-binding site presents a striking parallel to dimeric complexes of viral RNAs, which use the same hexanucleotide sequence for complexation and feature a similarly positioned metal ion-binding site within the dimeric structure. Taken together, these results indicate that the unique dimeric complex formed by the hs mt tRNALeuUUR A3243G mutant exhibits interesting similarities to biological RNA dimers, and may play a role in the loss of function caused by this mutation in vivo.
Figures
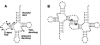


Similar articles
-
Dimerization of a pathogenic human mitochondrial tRNA.Nat Struct Biol. 2002 Aug;9(8):586-90. doi: 10.1038/nsb820. Nat Struct Biol. 2002. PMID: 12101407
-
The pathogenic U3271C human mitochondrial tRNA(Leu(UUR)) mutation disrupts a fragile anticodon stem.Nucleic Acids Res. 2003 Jan 15;31(2):596-601. doi: 10.1093/nar/gkg131. Nucleic Acids Res. 2003. PMID: 12527767 Free PMC article.
-
Combinatorial analysis of loop nucleotides in human mitochondrial tRNALeu(UUR).Biochemistry. 2005 Jan 11;44(1):233-42. doi: 10.1021/bi0489560. Biochemistry. 2005. PMID: 15628864
-
Dimer formation by tRNAs.Biochemistry (Mosc). 1999 Mar;64(3):298-306. Biochemistry (Mosc). 1999. PMID: 10205299 Review.
-
Probing RNA structure and metal-binding sites using terbium(III) footprinting.Curr Protoc Nucleic Acid Chem. 2003 Aug;Chapter 6:Unit 6.8. doi: 10.1002/0471142700.nc0608s13. Curr Protoc Nucleic Acid Chem. 2003. PMID: 18428913 Review.
Cited by
-
Structural Insights into RNA Dimerization: Motifs, Interfaces and Functions.Molecules. 2020 Jun 23;25(12):2881. doi: 10.3390/molecules25122881. Molecules. 2020. PMID: 32585844 Free PMC article. Review.
-
Native tertiary structure and nucleoside modifications suppress tRNA's intrinsic ability to activate the innate immune sensor PKR.PLoS One. 2013;8(3):e57905. doi: 10.1371/journal.pone.0057905. Epub 2013 Mar 4. PLoS One. 2013. PMID: 23483938 Free PMC article.
-
Global mapping of RNA homodimers in living cells.Genome Res. 2022 May;32(5):956-967. doi: 10.1101/gr.275900.121. Epub 2022 Mar 24. Genome Res. 2022. PMID: 35332098 Free PMC article.
-
The concept of RNA-assisted protein folding: the role of tRNA.Theor Biol Med Model. 2012 Apr 2;9:10. doi: 10.1186/1742-4682-9-10. Theor Biol Med Model. 2012. PMID: 22462735 Free PMC article.
-
Post-transcriptional nucleotide modification and alternative folding of RNA.Nucleic Acids Res. 2006 Feb 1;34(2):721-33. doi: 10.1093/nar/gkj471. Print 2006. Nucleic Acids Res. 2006. PMID: 16452298 Free PMC article. Review.
References
-
- Behlen, L.S., Sampson, J.R., DiRenzo, A.B., and Uhlenbeck, O.C. 1990. Lead-catalyzed cleavage of yeast tRNAPhe mutants. Biochemistry 29: 2515–2523. - PubMed
-
- Borner, G.V., Zeviani, M., Tiranti, V., Carrara, F., Hoffmann, S., Gerbitz, K.D., Lochmuller, H., Pongratz, D., Klopstock, T., Melberg, A., et al. 2000. Decreased aminoacylation of mutant tRNAs in MELAS but not in MERRF patients. Hum. Mol. Genet. 9: 467–475. - PubMed
-
- Chomyn, A., Martinuzzi, A., Yoneda, M., Daga, A., Hurko, O., Johns, D., Lai, S.T., Nonaka, I., Angelini, C., and Attardi, G. 1992. MELAS mutation in mtDNA binding site for transcription termination factor causes defects in protein synthesis and in respiration but no change in levels of upstream and downstream mature transcripts. Proc. Natl. Acad. Sci. 89: 4221–4225. - PMC - PubMed
-
- Chomyn, A., Enriquez, J.A., Micol, V., Fernandez-Silva, P., and Attardi, G. 2000. The mitochondrial myopathy, encephalopathy, lactic acidosis, and stroke-like episode syndrome-associated human mitochondrial tRNALeu(UUR) mutation causes aminoacylation deficiency and concomitant reduced association of mRNA with ribosomes. J. Biol. Chem. 275: 19198–19209. - PubMed
-
- El Meziane, A., Lehtinen, S.K., Holt, I.J., and Jacobs, H.T. 1998. Mitochondrial tRNALeu isoforms in lung carcinoma cybrid cells containing the np 3243 mtDNA mutation. Hum. Mol. Genet. 7: 2141–2147. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
