A preformed compact ribosome-binding domain in the cricket paralysis-like virus IRES RNAs
- PMID: 15701733
- PMCID: PMC1370722
- DOI: 10.1261/rna.7184705
A preformed compact ribosome-binding domain in the cricket paralysis-like virus IRES RNAs
Abstract
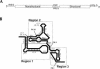
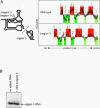
The internal ribosome site RNA of the cricket paralysis-like viruses (CrPV-like) binds directly to the ribosome, assembling the translation machinery without initiation factors. This mechanism does not require initiator tRNA, and translation starts from a non-AUG codon. A wealth of biochemical data has yielded a working model for this process, but the three-dimensional structure and biophysical characteristics of the unbound CrPV-like IRES RNAs are largely unexplored. Here, we demonstrate that the CrPV-like IRESes prefold into a two-part structure in the presence of magnesium ions. The largest part is a prefolded compact RNA domain that shares folding and structural characteristics with other compactly folded RNAs such as group I intron RNAs and RNase P RNA. Chemical probing reveals that the CrPV-like IRES' compact domain contains RNA helices that are packed tightly enough to exclude solvent, and analytical ultracentrifugation indicates a large change in the shape of the IRES upon folding. Formation of this compact domain is necessary for binding of the 40S subunit, and the structural organization of the unbound IRES RNA is consistent with the hypothesis that the IRES is functionally and structurally preorganized before ribosome binding.
Figures




References
-
- Adams, P.L., Stahley, M.R., Kosek, A.B., Wang, J., and Strobel, S.A. 2004. Crystal structure of a self-splicing group I intron with both exons. Nature 430: 45–50. - PubMed
-
- Cate, J.H., Gooding, A.R., Podell, E., Zhou, K., Golden, B.L., Kundrot, C.E., Cech, T.R., and Doudna, J.A. 1996. Crystal structure of a group I ribozyme domain: Principles of RNA packing. Science 273: 1678–1685. - PubMed
-
- Celander, D.W. and Cech, T.R. 1991. Visualizing the higher order folding of a catalytic RNA molecule. Science 251: 401–407. - PubMed
-
- Christian, P.D. and Scotti, P.D. 1998. Picornaviruses of insects. In The insect viruses (eds. L.K. Miller and A. Ball), pp. 301–336. Plenum, New York.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
