Saccharomyces cerevisiae imports the cytosolic pathway for Gln-tRNA synthesis into the mitochondrion
- PMID: 15706032
- PMCID: PMC551578
- DOI: 10.1101/gad.1269305
Saccharomyces cerevisiae imports the cytosolic pathway for Gln-tRNA synthesis into the mitochondrion
Abstract
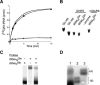
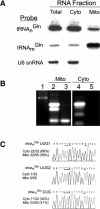
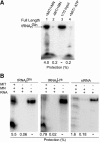
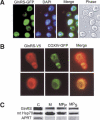
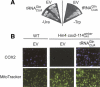
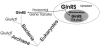
Aminoacyl-tRNA (aa-tRNA) formation, an essential process in protein biosynthesis, is generally achieved by direct attachment of an amino acid to tRNA by the aa-tRNA synthetases. An exception is Gln-tRNA synthesis, which in eukaryotes is catalyzed by glutaminyl-tRNA synthetase (GlnRS), while most bacteria, archaea, and chloroplasts employ the transamidation pathway, in which a tRNA-dependent glutamate modification generates Gln-tRNA. Mitochondrial protein synthesis is carried out normally by mitochondrial enzymes and organelle-encoded tRNAs that are different from their cytoplasmic counterparts. Early work suggested that mitochondria use the transamidation pathway for Gln-tRNA formation. We found no biochemical support for this in Saccharomyces cerevisiae mitochondria, but demonstrated the presence of the cytoplasmic GlnRS in the organelle and its involvement in mitochondrial Gln-tRNA synthesis. In addition, we showed in vivo localization of cytoplasmic tRNAGln in mitochondria and demonstrated its role in mitochondrial translation. We furthermore reconstituted in vitro cytoplasmic tRNAGln import into mitochondria by a novel mechanism. This tRNA import mechanism expands our knowledge of RNA trafficking in the eukaryotic cell. These findings change our view of the evolution of organellar protein synthesis.
Figures
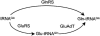
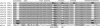
Similar articles
-
Trans-kingdom rescue of Gln-tRNAGln synthesis in yeast cytoplasm and mitochondria.Nucleic Acids Res. 2012 Oct;40(18):9171-81. doi: 10.1093/nar/gks689. Epub 2012 Jul 20. Nucleic Acids Res. 2012. PMID: 22821561 Free PMC article.
-
Widespread use of the glu-tRNAGln transamidation pathway among bacteria. A member of the alpha purple bacteria lacks glutaminyl-trna synthetase.J Biol Chem. 1996 Jun 21;271(25):14856-63. doi: 10.1074/jbc.271.25.14856. J Biol Chem. 1996. PMID: 8662929
-
Yeast mitochondrial Gln-tRNA(Gln) is generated by a GatFAB-mediated transamidation pathway involving Arc1p-controlled subcellular sorting of cytosolic GluRS.Genes Dev. 2009 May 1;23(9):1119-30. doi: 10.1101/gad.518109. Genes Dev. 2009. PMID: 19417106 Free PMC article.
-
Divergence of glutamate and glutamine aminoacylation pathways: providing the evolutionary rationale for mischarging.J Mol Evol. 1995 May;40(5):476-81. doi: 10.1007/BF00166615. J Mol Evol. 1995. PMID: 7783222 Review.
-
Mitochondrial tRNA import and its consequences for mitochondrial translation.Annu Rev Biochem. 2011;80:1033-53. doi: 10.1146/annurev-biochem-060109-092838. Annu Rev Biochem. 2011. PMID: 21417719 Review.
Cited by
-
Maf1 is involved in coupling carbon metabolism to RNA polymerase III transcription.Mol Cell Biol. 2007 Nov;27(21):7693-702. doi: 10.1128/MCB.01051-07. Epub 2007 Sep 4. Mol Cell Biol. 2007. PMID: 17785443 Free PMC article.
-
Role of the translocase of the mitochondrial inner membrane in the import of tRNAs into mitochondria in Trypanosoma brucei.Gene. 2020 Jul 20;748:144705. doi: 10.1016/j.gene.2020.144705. Epub 2020 Apr 24. Gene. 2020. PMID: 32339625 Free PMC article.
-
Mitoepigenetics and Its Emerging Roles in Cancer.Front Cell Dev Biol. 2020 Jan 23;8:4. doi: 10.3389/fcell.2020.00004. eCollection 2020. Front Cell Dev Biol. 2020. PMID: 32039210 Free PMC article. Review.
-
Trans-kingdom rescue of Gln-tRNAGln synthesis in yeast cytoplasm and mitochondria.Nucleic Acids Res. 2012 Oct;40(18):9171-81. doi: 10.1093/nar/gks689. Epub 2012 Jul 20. Nucleic Acids Res. 2012. PMID: 22821561 Free PMC article.
-
RNA-dependent conversion of phosphoserine forms selenocysteine in eukaryotes and archaea.Proc Natl Acad Sci U S A. 2006 Dec 12;103(50):18923-7. doi: 10.1073/pnas.0609703104. Epub 2006 Dec 1. Proc Natl Acad Sci U S A. 2006. PMID: 17142313 Free PMC article.
References
-
- Ahel I., Stathopoulos, C., Ambrogelly, A., Sauerwald, A., Toogood, H., Hartsch, T., and Söll, D. 2002. Cysteine activation is an inherent in vitro property of prolyl-tRNA synthetases. J. Biol. Chem. 277: 34743-34748. - PubMed
-
- Brown J.R. and Doolittle, W.F. 1999. Gene descent, duplication, and horizontal transfer in the evolution of glutamyl- and glutaminyl-tRNA synthetases. J. Mol. Evol. 49: 485-495. - PubMed
-
- Celis J.E. 1994. Cell biology: A laboratory handbook. Academic Press, San Diego, CA.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
