An acidic protein, YBAP1, mediates the release of YB-1 from mRNA and relieves the translational repression activity of YB-1
- PMID: 15713634
- PMCID: PMC549371
- DOI: 10.1128/MCB.25.5.1779-1792.2005
An acidic protein, YBAP1, mediates the release of YB-1 from mRNA and relieves the translational repression activity of YB-1
Abstract
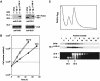
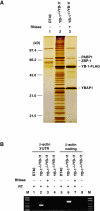
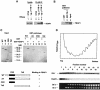
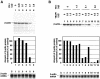
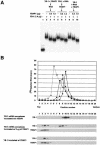
Eukaryotic Y-box proteins are nucleic acid-binding proteins implicated in a wide range of gene regulatory mechanisms. They contain the cold shock domain, which is a nucleic acid-binding structure also found in bacterial cold shock proteins. The Y-box protein YB-1 is known to be a core component of messenger ribonucleoprotein particles (mRNPs) in the cytoplasm. Here we disrupted the YB-1 gene in chicken DT40 cells. Through the immunoprecipitation of an epitope-tagged YB-1 protein, which complemented the slow-growth phenotype of YB-1-depleted cells, we isolated YB-1-associated complexes that likely represented general mRNPs in somatic cells. RNase treatment prior to immunoprecipitation led to the identification of a Y-box protein-associated acidic protein (YBAP1). The specific association of YB-1 with YBAP1 resulted in the release of YB-1 from reconstituted YB-1-mRNA complexes, thereby reducing the translational repression caused by YB-1 in the in vitro system. Our data suggest that YBAP1 induces the remodeling of YB-1-mRNA complexes.
Figures

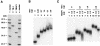

References
-
- Akey, C. W., and K. Luger. 2003. Histone chaperones and nucleosome assembly. Curr. Opin. Struct. Biol. 13:6-14. - PubMed
-
- Aoki, K., K. Matsumoto, and M. Tsujimoto. 2003. Xenopus cold-inducible RNA-binding protein 2 interacts with ElrA, the Xenopus homolog of HuR, and inhibits deadenylation of specific mRNAs. J. Biol. Chem. 278:48491-48497. - PubMed
-
- Bargou, R. C., K. Jurchott, C. Wagener, S. Bergmann, S. Metzner, K. Bommert, M. Y. Mapara, K. J. Winzer, M. Dietel, B. Dorken, and H. D. Royer. 1997. Nuclear localization and increased levels of transcription factor YB-1 in primary human breast cancers are associated with intrinsic MDR1 gene expression. Nat. Med. 3:447-450. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
