Comparisons of maize pericarp color1 alleles reveal paralogous gene recombination and an organ-specific enhancer region
- PMID: 15722466
- PMCID: PMC1069707
- DOI: 10.1105/tpc.104.029660
Comparisons of maize pericarp color1 alleles reveal paralogous gene recombination and an organ-specific enhancer region
Abstract
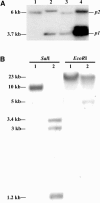
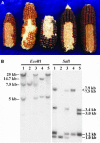
The maize (Zea mays) p1 (for pericarp color1) gene encodes an R2R3 Myb-like transcription factor that regulates the flavonoid biosynthetic pathway in floral organs, most notably kernel pericarp and cob. Alleles of the p1 gene condition distinct tissue-specific pigmentation patterns; to elucidate the molecular basis of these allele-specific expression patterns, we characterized two novel P1-rw (for red pericarp/white cob) alleles, P1-rw1077 and P1-rw751Ac. Structural analysis of P1-rw1077 indicated that this allele was generated by recombination between p1 and the tightly linked paralogous gene, p2. In the resulting gene, the p1 coding sequence was replaced by the p2 coding sequence, whereas the flanking p1 regulatory sequences remained largely preserved. The red pericarp color specified by P1-rw1077 suggests that the p1- and p2-encoded proteins are functionally equivalent as regulatory factors in the flavonoid biosynthesis pathway. Sequence analysis shows that the P1-rw1077 allele lacks a 386-bp sequence in a distal enhancer region 5 kb upstream of the transcription start site. An independently derived P1-rw allele contains an Ac insertion into the same sequence, indicating that this site likely contains cob glume-specific regulatory elements.
Figures







Similar articles
-
Gene structure induced epigenetic modifications of pericarp color1 alleles of maize result in tissue-specific mosaicism.PLoS One. 2009 Dec 14;4(12):e8231. doi: 10.1371/journal.pone.0008231. PLoS One. 2009. PMID: 20011605 Free PMC article.
-
Gene conversion between direct noncoding repeats promotes genetic and phenotypic diversity at a regulatory locus of Zea mays (L.).Genetics. 2006 Oct;174(2):753-62. doi: 10.1534/genetics.105.053942. Epub 2006 Jul 2. Genetics. 2006. PMID: 16816430 Free PMC article.
-
Epigenetic modifications of distinct sequences of the p1 regulatory gene specify tissue-specific expression patterns in maize.Genetics. 2007 Mar;175(3):1059-70. doi: 10.1534/genetics.106.066134. Epub 2006 Dec 18. Genetics. 2007. PMID: 17179091 Free PMC article.
-
Progressive loss of DNA methylation releases epigenetic gene silencing from a tandemly repeated maize Myb gene.Genetics. 2009 Jan;181(1):81-91. doi: 10.1534/genetics.108.097170. Epub 2008 Nov 10. Genetics. 2009. PMID: 19001287 Free PMC article.
-
Long-distance cis and trans interactions mediate paramutation.Adv Genet. 2002;46:215-34. doi: 10.1016/s0065-2660(02)46008-7. Adv Genet. 2002. PMID: 11931225 Review. No abstract available.
Cited by
-
The R2R3-MYB transcription factor gene family in maize.PLoS One. 2012;7(6):e37463. doi: 10.1371/journal.pone.0037463. Epub 2012 Jun 7. PLoS One. 2012. PMID: 22719841 Free PMC article.
-
Gene structure induced epigenetic modifications of pericarp color1 alleles of maize result in tissue-specific mosaicism.PLoS One. 2009 Dec 14;4(12):e8231. doi: 10.1371/journal.pone.0008231. PLoS One. 2009. PMID: 20011605 Free PMC article.
-
Divergence of gene regulation through chromosomal rearrangements.BMC Genomics. 2010 Nov 30;11:678. doi: 10.1186/1471-2164-11-678. BMC Genomics. 2010. PMID: 21118519 Free PMC article.
-
Development of high-resolution multiple-SNP arrays for genetic analyses and molecular breeding through genotyping by target sequencing and liquid chip.Plant Commun. 2021 Aug 9;2(6):100230. doi: 10.1016/j.xplc.2021.100230. eCollection 2021 Nov 8. Plant Commun. 2021. PMID: 34778746 Free PMC article.
-
5' Regulatory region of ubiquitin 2 gene from Porteresia coarctata makes efficient promoters for transgene expression in monocots and dicots.Plant Cell Rep. 2013 Aug;32(8):1199-210. doi: 10.1007/s00299-013-1416-3. Epub 2013 Mar 19. Plant Cell Rep. 2013. PMID: 23508257
References
-
- Belmaaza, A., and Chartrand, P. (1994). One-sided invasion events in homologous recombination at double-strand breaks. Mutat. Res. 314, 199–208. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources

