Biochemical characterization and DNA repair pathway interactions of Mag1-mediated base excision repair in Schizosaccharomyces pombe
- PMID: 15722486
- PMCID: PMC549418
- DOI: 10.1093/nar/gki259
Biochemical characterization and DNA repair pathway interactions of Mag1-mediated base excision repair in Schizosaccharomyces pombe
Abstract
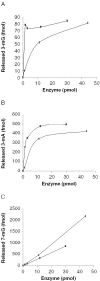
The Schizosaccharomyces pombe mag1 gene encodes a DNA repair enzyme with sequence similarity to the AlkA family of DNA glycosylases, which are essential for the removal of cytotoxic alkylation products, the premutagenic deamination product hypoxanthine and certain cyclic ethenoadducts such as ethenoadenine. In this paper, we have purified the Mag1 protein and characterized its substrate specificity. It appears that the substrate range of Mag1 is limited to the major alkylation products, such as 3-mA, 3-mG and 7-mG, whereas no significant activity was found towards deamination products, ethenoadducts or oxidation products. The efficiency of 3-mA and 3-mG removal was 5-10 times slower for Mag1 than for Escherichia coli AlkA whereas the rate of 7-mG removal was similar to the two enzymes. The relatively low efficiency for the removal of cytotoxic 3-methylpurines is consistent with the moderate sensitivity of the mag1 mutant to methylating agents. Furthermore, we studied the initial steps of Mag1-dependent base excision repair (BER) and genetic interactions with other repair pathways by mutant analysis. The double mutants mag1 nth1, mag1 apn2 and mag1 rad2 displayed increased resistance to methyl methanesulfonate (MMS) compared with the single mutants nth1, apn2 and rad2, respectively, indicating that Mag1 initiates both short-patch (Nth1-dependent) and long-patch (Rad2-dependent) BER of MMS-induced damage. Spontaneous intrachromosomal recombination frequencies increased 3-fold in the mag1 mutant suggesting that Mag1 and recombinational repair (RR) are both involved in repair of alkylated bases. Finally, we show that the deletion of mag1 in the background of rad16, nth1 and rad2 single mutants reduced the total recombination frequencies of all three double mutants, indicating that abasic sites formed as a result of Mag1 removal of spontaneous base lesions are substrates for nucleotide excision repair, long- and short-patch BER and RR.
Figures



Similar articles
-
Involvement of 3-methyladenine DNA glycosylases Mag1p and Mag2p in base excision repair of methyl methanesulfonate-damaged DNA in the fission yeast Schizosaccharomyces pombe.Genes Genet Syst. 2007 Dec;82(6):489-94. doi: 10.1266/ggs.82.489. Genes Genet Syst. 2007. PMID: 18270439
-
A general role of the DNA glycosylase Nth1 in the abasic sites cleavage step of base excision repair in Schizosaccharomyces pombe.Nucleic Acids Res. 2004 Sep 27;32(17):5119-25. doi: 10.1093/nar/gkh851. Print 2004. Nucleic Acids Res. 2004. PMID: 15452279 Free PMC article.
-
Roles of base excision repair enzymes Nth1p and Apn2p from Schizosaccharomyces pombe in processing alkylation and oxidative DNA damage.DNA Repair (Amst). 2005 Nov 21;4(11):1270-80. doi: 10.1016/j.dnarep.2005.06.009. Epub 2005 Aug 1. DNA Repair (Amst). 2005. PMID: 16076563
-
Mammalian DNA beta-polymerase in base excision repair of alkylation damage.Prog Nucleic Acid Res Mol Biol. 2001;68:57-74. doi: 10.1016/s0079-6603(01)68090-5. Prog Nucleic Acid Res Mol Biol. 2001. PMID: 11554313 Review.
-
Repair of oxidative DNA damage: mechanisms and functions.Cell Biochem Biophys. 2001;35(2):141-70. doi: 10.1385/CBB:35:2:141. Cell Biochem Biophys. 2001. PMID: 11892789 Review.
Cited by
-
Molecular characterization of Plasmodium falciparum DNA-3-methyladenine glycosylase.Malar J. 2020 Aug 6;19(1):284. doi: 10.1186/s12936-020-03355-w. Malar J. 2020. PMID: 32762689 Free PMC article.
-
Sculpting of DNA at abasic sites by DNA glycosylase homolog mag2.Structure. 2013 Jan 8;21(1):154-166. doi: 10.1016/j.str.2012.11.004. Epub 2012 Dec 13. Structure. 2013. PMID: 23245849 Free PMC article.
-
A New Family of HEAT-Like Repeat Proteins Lacking a Critical Substrate Recognition Motif Present in Related DNA Glycosylases.PLoS One. 2015 May 15;10(5):e0127733. doi: 10.1371/journal.pone.0127733. eCollection 2015. PLoS One. 2015. PMID: 25978435 Free PMC article.
-
Proficient repair in chromatin remodeling defective ino80 mutants of Saccharomyces cerevisiae highlights replication defects as the main contributor to DNA damage sensitivity.DNA Repair (Amst). 2010 Sep 4;9(9):976-84. doi: 10.1016/j.dnarep.2010.06.010. Epub 2010 Jul 31. DNA Repair (Amst). 2010. PMID: 20674516 Free PMC article.
-
Genetic Interaction Landscape Reveals Critical Requirements for Schizosaccharomyces pombe Brc1 in DNA Damage Response Mutants.G3 (Bethesda). 2015 Mar 19;5(5):953-62. doi: 10.1534/g3.115.017251. G3 (Bethesda). 2015. PMID: 25795664 Free PMC article.
References
-
- Dianov G.L., Sleeth K.M., Dianova I.I., Allinson S.L. Repair of abasic sites in DNA. Mutat. Res. 2003;531:157–163. - PubMed
-
- Slupphaug G., Kavli B., Krokan H.E. The interacting pathways for prevention and repair of oxidative DNA damage. Mutat. Res. 2003;531:231–251. - PubMed
-
- Falnes P.O., Johansen R.F., Seeberg E. AlkB-mediated oxidative demethylation reverses DNA damage in Escherichia coli. Nature. 2002;419:178–182. - PubMed
-
- Trewick S.C., Henshaw T.F., Hausinger R.P., Lindahl T., Sedgwick B. Oxidative demethylation by Escherichia coli AlkB directly reverts DNA base damage. Nature. 2002;419:174–178. - PubMed
-
- Laval J. Two enzymes are required from strand incision in repair of alkylated DNA. Nature. 1977;269:829–832. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

