Simocyclinone D8, an inhibitor of DNA gyrase with a novel mode of action
- PMID: 15728908
- PMCID: PMC549283
- DOI: 10.1128/AAC.49.3.1093-1100.2005
Simocyclinone D8, an inhibitor of DNA gyrase with a novel mode of action
Abstract
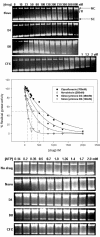
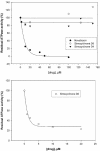
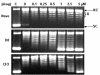
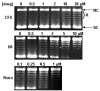
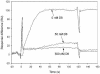
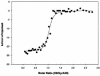
We have characterized the interaction of a new class of antibiotics, simocyclinones, with bacterial DNA gyrase. Even though their structures include an aminocoumarin moiety, a key feature of novobiocin, coumermycin A(1), and clorobiocin, which also target gyrase, simocyclinones behave strikingly differently from these compounds. Simocyclinone D8 is a potent inhibitor of gyrase supercoiling, with a 50% inhibitory concentration lower than that of novobiocin. However, it does not competitively inhibit the DNA-independent ATPase reaction of GyrB, which is characteristic of other aminocoumarins. Simocyclinone D8 also inhibits DNA relaxation by gyrase but does not stimulate cleavage complex formation, unlike quinolones, the other major class of gyrase inhibitors; instead, it abrogates both Ca(2+)- and quinolone-induced cleavage complex formation. Binding studies suggest that simocyclinone D8 interacts with the N-terminal domain of GyrA. Taken together, our results demonstrate that simocyclinones inhibit an early step of the gyrase catalytic cycle by preventing binding of the enzyme to DNA. This is a novel mechanism for a gyrase inhibitor and presents new possibilities for antibacterial drug development.
Figures


Similar articles
-
Mapping simocyclinone D8 interaction with DNA gyrase: evidence for a new binding site on GyrB.Antimicrob Agents Chemother. 2010 Jan;54(1):213-20. doi: 10.1128/AAC.00972-09. Epub 2009 Oct 26. Antimicrob Agents Chemother. 2010. PMID: 19858260 Free PMC article.
-
A crystal structure of the bifunctional antibiotic simocyclinone D8, bound to DNA gyrase.Science. 2009 Dec 4;326(5958):1415-8. doi: 10.1126/science.1179123. Science. 2009. PMID: 19965760
-
Inhibition of human topoisomerases I and II by simocyclinone D8.J Nat Prod. 2012 Aug 24;75(8):1485-9. doi: 10.1021/np300299y. Epub 2012 Aug 6. J Nat Prod. 2012. PMID: 22867097
-
New aminocoumarin antibiotics as gyrase inhibitors.Int J Med Microbiol. 2014 Jan;304(1):31-6. doi: 10.1016/j.ijmm.2013.08.013. Epub 2013 Sep 4. Int J Med Microbiol. 2014. PMID: 24079980 Review.
-
The interaction between coumarin drugs and DNA gyrase.Mol Microbiol. 1993 Aug;9(4):681-6. doi: 10.1111/j.1365-2958.1993.tb01728.x. Mol Microbiol. 1993. PMID: 8231802 Review.
Cited by
-
Avoidance of suicide in antibiotic-producing microbes.J Ind Microbiol Biotechnol. 2010 Jul;37(7):643-72. doi: 10.1007/s10295-010-0721-x. Epub 2010 May 6. J Ind Microbiol Biotechnol. 2010. PMID: 20446033 Review.
-
Molecular basis for the differential quinolone susceptibility of mycobacterial DNA gyrase.Antimicrob Agents Chemother. 2014;58(4):2013-20. doi: 10.1128/AAC.01958-13. Epub 2014 Jan 13. Antimicrob Agents Chemother. 2014. PMID: 24419347 Free PMC article.
-
The Macromolecular Machines that Duplicate the Escherichia coli Chromosome as Targets for Drug Discovery.Antibiotics (Basel). 2018 Mar 14;7(1):23. doi: 10.3390/antibiotics7010023. Antibiotics (Basel). 2018. PMID: 29538288 Free PMC article. Review.
-
s-Triazine Derivatives Functionalized with Alkylating 2-Chloroethylamine Fragments as Promising Antimicrobial Agents: Inhibition of Bacterial DNA Gyrases, Molecular Docking Studies, and Antibacterial and Antifungal Activity.Pharmaceuticals (Basel). 2023 Sep 4;16(9):1248. doi: 10.3390/ph16091248. Pharmaceuticals (Basel). 2023. PMID: 37765056 Free PMC article.
-
The naphthoquinone diospyrin is an inhibitor of DNA gyrase with a novel mechanism of action.J Biol Chem. 2013 Feb 15;288(7):5149-56. doi: 10.1074/jbc.M112.419069. Epub 2012 Dec 28. J Biol Chem. 2013. PMID: 23275348 Free PMC article.
References
-
- Abdelfattah, M., R. P. Maskey, R. N. Asolkar, I. Grun-Wollny, and H. Laatsch. 2003. Seitomycin: isolation, structure elucidation and biological activity of a new angucycline antibiotic from a terrestrial streptomycete. J. Antibiot. 56:539-542. - PubMed
-
- Ali, J. A., A. P. Jackson, A. J. Howells, and A. Maxwell. 1993. The 43-kDa N-terminal fragment of the gyrase B protein hydrolyses ATP and binds coumarin drugs. Biochemistry 32:2717-2724. - PubMed
-
- Bannister, B., and B. A. Zapotocky. 1992. Protorubradirin, an antibiotic containing a C-nitroso-sugar fragment, is the true secondary metabolite produced by Streptomyces achromogenes var. rubradiris. Rubradirin, described earlier, is its photo-oxidation product. J. Antibiot. 45:1313-1324. - PubMed
-
- Champoux, J. J. 2001. DNA topoisomerases: structure, function, and mechanism. Annu. Rev. Biochem. 70:369-413. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

