Altered gingipain maturation in vimA- and vimE-defective isogenic mutants of Porphyromonas gingivalis
- PMID: 15731033
- PMCID: PMC1064936
- DOI: 10.1128/IAI.73.3.1357-1366.2005
Altered gingipain maturation in vimA- and vimE-defective isogenic mutants of Porphyromonas gingivalis
Abstract
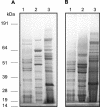
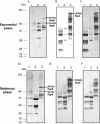
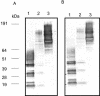
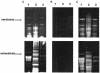
We have previously shown that gingipain activity in Porphyromonas gingivalis is modulated by the unique vimA and vimE genes. To determine if these genes had a similar phenotypic effect on protease maturation and activation, isogenic mutants defective in those genes were further characterized. Western blot analyses with antigingipain antibodies showed RgpA-, RgpB-, and Kgp-immunoreactive bands in membrane fractions as well as the culture supernatant of both P. gingivalis W83 and FLL93, the vimE-defective mutant. In contrast, the membrane of P. gingivalis FLL92, the vimA-defective mutant, demonstrated immunoreactivity only with RgpB antibodies. With mass spectrometry or Western blots, full-length RgpA and RgpB were identified from extracellular fractions. In similar extracellular fractions from P. gingivalis FLL92 and FLL93, purified RgpB activated only arginine-specific activity. In addition, the lipopolysaccharide profiles of the vimA and vimE mutants were truncated in comparison to that of W83. While glycosylated proteins were detected in the membrane and extracellular fractions from the vimA- and vimE-defective mutants, a monoclonal antibody (1B5) that reacts with specific sugar moieties of the P. gingivalis cell surface polysaccharide and membrane-associated Rgp gingipain showed no immunoreactivity with these fractions. Taken together, these results indicate a possible defect in sugar biogenesis in both the vimA- and vimE-defective mutants. These modulating genes play a role in the secretion, processing, and/or anchorage of gingipains on the cell surface.
Figures

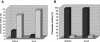




Similar articles
-
The vimE gene downstream of vimA is independently expressed and is involved in modulating proteolytic activity in Porphyromonas gingivalis W83.Infect Immun. 2004 Oct;72(10):5555-64. doi: 10.1128/IAI.72.10.5555-5564.2004. Infect Immun. 2004. PMID: 15385452 Free PMC article.
-
The roles of RgpB and Kgp in late onset gingipain activity in the vimA-defective mutant of Porphyromonas gingivalis W83.Mol Oral Microbiol. 2015 Oct;30(5):347-60. doi: 10.1111/omi.12098. Epub 2015 May 8. Mol Oral Microbiol. 2015. PMID: 25858089 Free PMC article.
-
Role of Acetyltransferase PG1842 in Gingipain Biogenesis in Porphyromonas gingivalis.J Bacteriol. 2018 Nov 26;200(24):e00385-18. doi: 10.1128/JB.00385-18. Print 2018 Dec 15. J Bacteriol. 2018. PMID: 30249709 Free PMC article.
-
Porphyromonas gingivalis gingipains: the molecular teeth of a microbial vampire.Curr Protein Pept Sci. 2003 Dec;4(6):409-26. doi: 10.2174/1389203033487009. Curr Protein Pept Sci. 2003. PMID: 14683427 Review.
-
Gingipains, the major cysteine proteinases and virulence factors of Porphyromonas gingivalis: structure, function and assembly of multidomain protein complexes.Curr Protein Pept Sci. 2003 Dec;4(6):397-407. doi: 10.2174/1389203033487036. Curr Protein Pept Sci. 2003. PMID: 14683426 Review.
Cited by
-
The bcp gene in the bcp-recA-vimA-vimE-vimF operon is important in oxidative stress resistance in Porphyromonas gingivalis W83.Mol Oral Microbiol. 2011 Feb;26(1):62-77. doi: 10.1111/j.2041-1014.2010.00596.x. Epub 2010 Nov 18. Mol Oral Microbiol. 2011. PMID: 21214873 Free PMC article.
-
Por secretion system-dependent secretion and glycosylation of Porphyromonas gingivalis hemin-binding protein 35.PLoS One. 2011;6(6):e21372. doi: 10.1371/journal.pone.0021372. Epub 2011 Jun 22. PLoS One. 2011. PMID: 21731719 Free PMC article.
-
Gingipain-dependent interactions with the host are important for survival of Porphyromonas gingivalis.Front Biosci. 2008 May 1;13:3215-38. doi: 10.2741/2922. Front Biosci. 2008. PMID: 18508429 Free PMC article. Review.
-
The native 67-kilodalton minor fimbria of Porphyromonas gingivalis is a novel glycoprotein with DC-SIGN-targeting motifs.J Bacteriol. 2010 Aug;192(16):4103-10. doi: 10.1128/JB.00275-10. Epub 2010 Jun 18. J Bacteriol. 2010. PMID: 20562309 Free PMC article.
-
The sweet tooth of bacteria: common themes in bacterial glycoconjugates.Microbiol Mol Biol Rev. 2014 Sep;78(3):372-417. doi: 10.1128/MMBR.00007-14. Microbiol Mol Biol Rev. 2014. PMID: 25184559 Free PMC article. Review.
References
-
- Abaibou, H., Q. Ma, G. J. Olango, J. Potempa, J. Travis, and H. M. Fletcher. 2000. Unaltered expression of the major protease genes in a non-virulent recA-defective mutant of Porphyromonas gingivalis W83. Oral Microbiol. Immunol. 15:40-47. - PubMed
-
- Aduse-Opoku, J., N. N. Davies, A. Gallagher, A. Hashim, H. E. Evans, M. Rangarajan, J. M. Slaney, and M. A. Curtis. 2000. Generation of lys-gingipain protease activity in Porphyromonas gingivalis W50 is independent of Arg-gingipain protease activities. Microbiology 146:1933-1940. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

