The first steps of transposable elements invasion: parasitic strategy vs. genetic drift
- PMID: 15731520
- PMCID: PMC1449084
- DOI: 10.1534/genetics.104.031211
The first steps of transposable elements invasion: parasitic strategy vs. genetic drift
Abstract
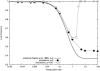
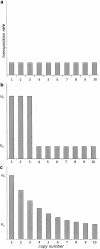
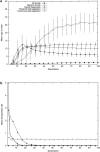
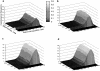
Transposable elements are often considered as selfish DNA sequences able to invade the genome of their host species. Their evolutive dynamics are complex, due to the interaction between their intrinsic amplification capacity, selection at the host level, transposition regulation, and genetic drift. Here, we propose modeling the first steps of TE invasion, i.e., just after a horizontal transfer, when a single copy is present in the genome of one individual. If the element has a constant transposition rate, it will disappear in most cases: the elements with low-transposition rate are frequently lost through genetic drift, while those with high-transposition rate may amplify, leading to the sterility of their host. Elements whose transposition rate is regulated are able to successfully invade the populations, thanks to an initial transposition burst followed by a strong limitation of their activity. Self-regulation or hybrid dysgenesis may thus represent some genome-invasion parasitic strategies.
Figures

References
-
- Anxolabéhère, D., M. G. Kidwell and G. Periquet, 1988. Molecular characteristics of diverse populations are consistent with the hypothesis of a recent invasion of Drosophila melanogaster by mobile P elements. Mol. Biol. Evol. 5: 252–269. - PubMed
-
- Biémont, C., 1994. Dynamic equilibrium between insertion and excision of P elements in highly inbred lines from an M′ strain of Drosophila melanogaster. J. Mol. Evol. 39: 466–472. - PubMed
-
- Biémont, C., F. Lemeunier, M. P. Garcia Guerreiro, J. F. Brookfield, C. Gautier et al., 1994. Population dynamics of the copia, mdg1, mdg3, gypsy, and P transposable elements in a natural population of Drosophila melanogaster. Genet. Res. 63: 197–212. - PubMed
-
- Biémont, C., C. Nardon, G. Deceliere, D. Lepetit, C. Loevenbruck et al., 2003. Worldwide distribution of transposable element copy number in natural populations of Drosophila simulans. Evolution 57: 159–167. - PubMed
-
- Bregliano, J.-C., and M. G. Kidwell, 1983 Hybrid dysgenesis determinants, p. 363 in Mobile Genetic Elements, edited by J. A. Shapiro. Academic Press, San Diego.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

