Independence of replisomes in Escherichia coli chromosomal replication
- PMID: 15738384
- PMCID: PMC552787
- DOI: 10.1073/pnas.0500812102
Independence of replisomes in Escherichia coli chromosomal replication
Abstract
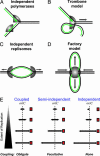
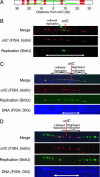
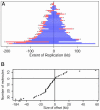
In Escherichia coli DNA replication is carried out by the coordinated action of the proteins within a replisome. After replication initiation, the two bidirectionally oriented replisomes from a single origin are colocalized into higher-order structures termed replication factories. The factory model postulated that the two replisomes are also functionally coupled. We tested this hypothesis by using DNA combing and whole-genome microarrays. Nascent DNA surrounding oriC in single, combed chromosomes showed instead that one replisome, usually the leftward one, was significantly ahead of the other 70% of the time. We next used microarrays to follow replication throughout the genome by measuring DNA copy number. We found in multiple E. coli strains that the replisomes are independent, with the leftward replisome ahead of the rightward one. The size of the bias was strain-specific, varying from 50 to 130 kb in the array results. When we artificially blocked one replisome, the other continued unabated, again demonstrating independence. We suggest an improved version of the factory model that retains the advantages of threading DNA through colocalized replisomes at about equal rates, but allows the cell flexibility to overcome obstacles encountered during elongation.
Figures



References
-
- Kornberg, A. & Baker, T. A. (1992) DNA Replication (Freeman, New York).
-
- Alberts, B. M., Barry, J., Bedinger, P., Formosa, T., Jongeneel, C. V. & Kreuzer, K. N. (1983) Cold Spring Harb. Symp. Quant. Biol. 47, 655–668. - PubMed
-
- Lee, J., Chastain, P. D., 2nd, Kusakabe, T., Griffith, J. D. & Richardson, C. C. (1998) Mol. Cell 1, 1001–1010. - PubMed
-
- Pages, V. & Fuchs, R. P. (2003) Science 300, 1300–1303. - PubMed
-
- Higuchi, K., Katayama, T., Iwai, S., Hidaka, M., Horiuchi, T. & Maki, H. (2003) Genes Cells 8, 437–449. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

