Long-range electron transfer
- PMID: 15738403
- PMCID: PMC553296
- DOI: 10.1073/pnas.0408029102
Long-range electron transfer
Abstract
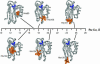
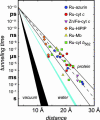
Recent investigations have shed much light on the nuclear and electronic factors that control the rates of long-range electron tunneling through molecules in aqueous and organic glasses as well as through bonds in donor-bridge-acceptor complexes. Couplings through covalent and hydrogen bonds are much stronger than those across van der Waals gaps, and these differences in coupling between bonded and nonbonded atoms account for the dependence of tunneling rates on the structure of the media between redox sites in Ru-modified proteins and protein-protein complexes.
Figures

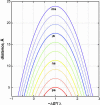

 ;H
;H P) with the indicated free-energy changes for the R
P) with the indicated free-energy changes for the R H step.
H step.References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

