Soilborne wheat mosaic virus (SBWMV) 19K protein belongs to a class of cysteine rich proteins that suppress RNA silencing
- PMID: 15740624
- PMCID: PMC555535
- DOI: 10.1186/1743-422X-2-18
Soilborne wheat mosaic virus (SBWMV) 19K protein belongs to a class of cysteine rich proteins that suppress RNA silencing
Abstract
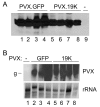
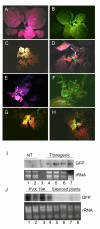
Amino acid sequence analyses indicate that the Soilborne wheat mosaic virus (SBWMV) 19K protein is a cysteine-rich protein (CRP) and shares sequence homology with CRPs derived from furo-, hordei-, peclu- and tobraviruses. Since the hordei- and pecluvirus CRPs were shown to be pathogenesis factors and/or suppressors of RNA silencing, experiments were conducted to determine if the SBWMV 19K CRP has similar activities. The SBWMV 19K CRP was introduced into the Potato virus X (PVX) viral vector and inoculated to tobacco plants. The SBWMV 19K CRP aggravated PVX-induced symptoms and restored green fluorescent protein (GFP) expression to GFP silenced tissues. These observations indicate that the SBWMV 19K CRP is a pathogenicity determinant and a suppressor of RNA silencing.
Figures


References
-
- Li WX, Li H, Lu R, Li F, Dus M, Atkinson P, Brydon EW, Johnson KL, Garcia-Sastre A, Ball LA, Palese P, Ding SW. Interferon antagonist proteins of influenza and vaccinia viruses are suppressors of RNA silencing. Proc Natl Acad Sci U S A. 2004;101:1350–1355. doi: 10.1073/pnas.0308308100. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

