Chemotaxonomic identification of single bacteria by micro-Raman spectroscopy: application to clean-room-relevant biological contaminations
- PMID: 15746368
- PMCID: PMC1065155
- DOI: 10.1128/AEM.71.3.1626-1637.2005
Chemotaxonomic identification of single bacteria by micro-Raman spectroscopy: application to clean-room-relevant biological contaminations
Abstract
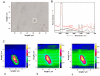
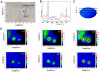
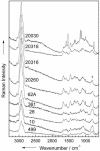
Microorganisms, such as bacteria, which might be present as contamination inside an industrial food or pharmaceutical clean room process need to be identified on short time scales in order to minimize possible health hazards as well as production downtimes causing financial deficits. Here we describe the first results of single-particle micro-Raman measurements in combination with a classification method, the so-called support vector machine technique, allowing for a fast, reliable, and nondestructive online identification method for single bacteria.
Figures





Similar articles
-
Bacillus spore classification via surface-enhanced Raman spectroscopy and principal component analysis.Appl Spectrosc. 2008 Mar;62(3):267-72. doi: 10.1366/000370208783759623. Appl Spectrosc. 2008. PMID: 18339232
-
Micro-Raman spectroscopic identification of bacterial cells of the genus Staphylococcus and dependence on their cultivation conditions.Analyst. 2005 Nov;130(11):1543-50. doi: 10.1039/b507715j. Epub 2005 Sep 30. Analyst. 2005. PMID: 16222378
-
Single bacteria identification by Raman spectroscopy.J Biomed Opt. 2014;19(11):111610. doi: 10.1117/1.JBO.19.11.111610. J Biomed Opt. 2014. PMID: 25028774
-
Characterisation and identification of bacteria using SERS.Chem Soc Rev. 2008 May;37(5):931-6. doi: 10.1039/b705973f. Epub 2008 Mar 26. Chem Soc Rev. 2008. PMID: 18443678 Review.
-
Vibrational spectroscopy--a powerful tool for the rapid identification of microbial cells at the single-cell level.Cytometry A. 2009 Feb;75(2):104-13. doi: 10.1002/cyto.a.20682. Cytometry A. 2009. PMID: 19156822 Review.
Cited by
-
Comprehensive detection and discrimination of Campylobacter species by use of confocal micro-Raman spectroscopy and multilocus sequence typing.J Clin Microbiol. 2012 Sep;50(9):2932-46. doi: 10.1128/JCM.01144-12. Epub 2012 Jun 27. J Clin Microbiol. 2012. PMID: 22740711 Free PMC article.
-
Raman spectroscopy as a potential tool for detection of Brucella spp. in milk.Appl Environ Microbiol. 2012 Aug;78(16):5575-83. doi: 10.1128/AEM.00637-12. Epub 2012 Jun 1. Appl Environ Microbiol. 2012. PMID: 22660699 Free PMC article.
-
Evaluation of the impact of buffered peptone water composition on the discrimination between Salmonella enterica and Escherichia coli by Raman spectroscopy.Anal Bioanal Chem. 2020 Jun;412(15):3595-3604. doi: 10.1007/s00216-020-02596-7. Epub 2020 Apr 4. Anal Bioanal Chem. 2020. PMID: 32248395
-
Microbial phenomics linking the phenotype to fonction: The potential of Raman spectroscopy.J Microbiol. 2021 Mar;59(3):249-258. doi: 10.1007/s12275-021-0590-1. Epub 2021 Jan 26. J Microbiol. 2021. PMID: 33496936 Review.
-
An ensemble approach to the structure-function problem in microbial communities.iScience. 2022 Jan 11;25(2):103761. doi: 10.1016/j.isci.2022.103761. eCollection 2022 Feb 18. iScience. 2022. PMID: 35141504 Free PMC article. Review.
References
-
- Alexander, T. A., P. M. Pellegrino, and J. B. Gillespie. 2003. Near-infrared surface-enhanced-Raman-scattering (SERS) mediated identification of single optically trapped bacterial spores. Proc. SPIE 5085:91-100. - PubMed
-
- Al-Khaldi, S. F., and M. M. Mossoba. 2004. Gene and bacterial identification using high-throughput technologies: genomics, proteomics, and phenomics. Nutrition 20:32-38. - PubMed
-
- Berger, A. J., and Q. Zhu. 2003. Identification of oral bacteria by Raman microspectroscopy. J. Mod. Optics 50:2375-2380.
-
- Britton, K. A., R. A. Dalterio, W. H. Nelson, D. Britt, and J. F. Sperry. 1988. Ultraviolet resonance Raman spectra of Escherichia coli with 222.5-251.0 nm pulsed laser excitation. Appl. Spectrosc. 42:782-788.
-
- Burges, C. J. C. 1998. A tutorial on support vector machines for pattern recognition. Data Mining Knowledge Disc. 2:121-167.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

