Nosocomial outbreak caused by Salmonella enterica serotype Livingstone producing CTX-M-27 extended-spectrum beta-lactamase in a neonatal unit in Sousse, Tunisia
- PMID: 15750057
- PMCID: PMC1081247
- DOI: 10.1128/JCM.43.3.1037-1044.2005
Nosocomial outbreak caused by Salmonella enterica serotype Livingstone producing CTX-M-27 extended-spectrum beta-lactamase in a neonatal unit in Sousse, Tunisia
Abstract
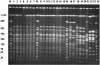
In this study, we report an outbreak of Salmonella enterica serotype Livingstone resistant to extended-spectrum cephalosporins that occurred in a neonatal ward of the maternity department of Farhat Hached Hospital, Sousse, Tunisia, in 2002. A total of 16 isolates were recovered from 16 babies hospitalized in the ward during the period 1 to 16 July. All these babies developed diarrhea, and three of them developed septicemia. All the isolates demonstrated resistance to ceftriaxone and ceftazidime due to the production of an extended-spectrum beta-lactamase (ESBL). The isolates were also resistant to aminoglycosides (kanamycin, tobramycin, netilmicin, gentamicin, and amikacin) and sulfamethoxazole-trimethoprim. DNA profiles were determined by pulsed-field gel electrophoresis using the XbaI and SpeI endonucleases and by ribotyping with PstI digestion. They yielded the same patterns, showing that the outbreak was caused by a single clone. The ESBL was identified as CTX-M-27 by sequencing of PCR products and by isoelectric focusing. The ESBL resistance was transferred by a 40-kb conjugative plasmid. The mobile insertion sequence ISEcp1 was found to be located upstream of bla(CTX-M-27) in the same position as that known for a bla(CTX-M-14) sequence. A new gene named dfrA21, encoding resistance to trimethoprim and carried by a 90-kb plasmid, was characterized. The dfrA21 gene was inserted as a single resistance cassette in a class I integron. The babies were treated with colistin, and all except two recovered. The outbreak came to an end when appropriate actions were taken: patient isolation, hand washing, and disinfection of the ward.
Figures


Similar articles
-
Prevalence and characterization of extended-spectrum beta-lactamases-producing Salmonella enterica isolates in Saragossa, Spain (2001-2008).Microb Drug Resist. 2011 Jun;17(2):207-13. doi: 10.1089/mdr.2010.0105. Epub 2011 Jan 16. Microb Drug Resist. 2011. PMID: 21235396
-
Multiple outbreaks of nosocomial salmonellosis in Russia and Belarus caused by a single clone of Salmonella enterica serovar Typhimurium producing an extended-spectrum beta-lactamase.Antimicrob Agents Chemother. 2004 Aug;48(8):2808-15. doi: 10.1128/AAC.48.8.2808-2815.2004. Antimicrob Agents Chemother. 2004. PMID: 15273085 Free PMC article.
-
Extended-spectrum {beta}-lactamases and AmpC {beta}-lactamases in ceftiofur-resistant Salmonella enterica isolates from food and livestock obtained in Germany during 2003-07.J Antimicrob Chemother. 2009 Aug;64(2):301-9. doi: 10.1093/jac/dkp195. Epub 2009 May 27. J Antimicrob Chemother. 2009. PMID: 19474065
-
Molecular epidemiology and genetic environment of acquired bla ACC-1 in Salmonella enterica serotype Livingstone causing a large nosocomial outbreak in Tunisia.Microb Drug Resist. 2009 Dec;15(4):279-86. doi: 10.1089/mdr.2009.0035. Microb Drug Resist. 2009. PMID: 19857134
-
An outbreak of extended-spectrum, beta-lactamase-producing Salmonella senftenberg in a burns ward.J Hosp Infect. 1998 Dec;40(4):295-302. doi: 10.1016/s0195-6701(98)90307-3. J Hosp Infect. 1998. PMID: 9868622 Review.
Cited by
-
Emergence and outbreaks of CTX-M beta-lactamase-producing Escherichia coli and Klebsiella pneumoniae strains in a Tunisian hospital.J Clin Microbiol. 2006 Nov;44(11):4049-56. doi: 10.1128/JCM.01076-06. Epub 2006 Sep 6. J Clin Microbiol. 2006. PMID: 16957046 Free PMC article.
-
CTX-M-3 and CTX-M-15 extended-spectrum beta-lactamases in isolates of Escherichia coli from a hospital in Algiers, Algeria.J Clin Microbiol. 2006 Dec;44(12):4584-6. doi: 10.1128/JCM.01445-06. Epub 2006 Sep 20. J Clin Microbiol. 2006. PMID: 16988017 Free PMC article.
-
Comparison of five molecular subtyping methods for differentiation of Salmonella Kentucky isolates in Tunisia.World J Microbiol Biotechnol. 2014 Jan;30(1):87-98. doi: 10.1007/s11274-013-1414-1. Epub 2013 Jul 10. World J Microbiol Biotechnol. 2014. PMID: 23839713
-
Chromosomal integration of the extended-spectrum beta-lactamase gene blaCTX-M-15 in Salmonella enterica serotype Concord isolates from internationally adopted children.Antimicrob Agents Chemother. 2009 May;53(5):1808-16. doi: 10.1128/AAC.00451-08. Epub 2009 Mar 9. Antimicrob Agents Chemother. 2009. PMID: 19273688 Free PMC article.
-
Dissemination and genetic support of broad-spectrum beta-lactam-resistant Escherichia coli strain isolated from two Tunisian hospitals during 2004-2012.Afr Health Sci. 2017 Jun;17(2):346-355. doi: 10.4314/ahs.v17i2.8. Afr Health Sci. 2017. PMID: 29062329 Free PMC article.
References
-
- Altschul, S. F., W. Gish, W. Miller, E. W. Myers, and D. J. Lipman. 1990. Basic local alignment search tool. J. Mol. Biol. 215:403-410. - PubMed
-
- Arlet, G., M. Rouveau, and A. Philippon. 1997. Substitution of alanine for aspartate at position 179 in the SHV-6 extended-spectrum β-lactamase. FEMS Microbiol. Lett. 152:163-167. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Medical

