Phylogenomic approaches to common problems encountered in the analysis of low copy repeats: the sulfotransferase 1A gene family example
- PMID: 15752422
- PMCID: PMC555591
- DOI: 10.1186/1471-2148-5-22
Phylogenomic approaches to common problems encountered in the analysis of low copy repeats: the sulfotransferase 1A gene family example
Abstract
Background: Blocks of duplicated genomic DNA sequence longer than 1000 base pairs are known as low copy repeats (LCRs). Identified by their sequence similarity, LCRs are abundant in the human genome, and are interesting because they may represent recent adaptive events, or potential future adaptive opportunities within the human lineage. Sequence analysis tools are needed, however, to decide whether these interpretations are likely, whether a particular set of LCRs represents nearly neutral drift creating junk DNA, or whether the appearance of LCRs reflects assembly error. Here we investigate an LCR family containing the sulfotransferase (SULT) 1A genes involved in drug metabolism, cancer, hormone regulation, and neurotransmitter biology as a first step for defining the problems that those tools must manage.
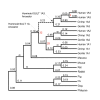
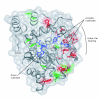
Results: Sequence analysis here identified a fourth sulfotransferase gene, which may be transcriptionally active, located on human chromosome 16. Four regions of genomic sequence containing the four human SULT1A paralogs defined a new LCR family. The stem hominoid SULT1A progenitor locus was identified by comparative genomics involving complete human and rodent genomes, and a draft chimpanzee genome. SULT1A expansion in hominoid genomes was followed by positive selection acting on specific protein sites. This episode of adaptive evolution appears to be responsible for the dopamine sulfonation function of some SULT enzymes. Each of the conclusions that this bioinformatic analysis generated using data that has uncertain reliability (such as that from the chimpanzee genome sequencing project) has been confirmed experimentally or by a "finished" chromosome 16 assembly, both of which were published after the submission of this manuscript.
Conclusion: SULT1A genes expanded from one to four copies in hominoids during intra-chromosomal LCR duplications, including (apparently) one after the divergence of chimpanzees and humans. Thus, LCRs may provide a means for amplifying genes (and other genetic elements) that are adaptively useful. Being located on and among LCRs, however, could make the human SULT1A genes susceptible to further duplications or deletions resulting in 'genomic diseases' for some individuals. Pharmacogenomic studies of SULT1Asingle nucleotide polymorphisms, therefore, should also consider examining SULT1A copy number variability when searching for genotype-phenotype associations. The latest duplication is, however, only a substantiated hypothesis; an alternative explanation, disfavored by the majority of evidence, is that the duplication is an artifact of incorrect genome assembly.
Figures


Similar articles
-
Duplication and relocation of the functional DPY19L2 gene within low copy repeats.BMC Genomics. 2006 Mar 9;7:45. doi: 10.1186/1471-2164-7-45. BMC Genomics. 2006. PMID: 16526957 Free PMC article.
-
Lineage-specific gene duplication and loss in human and great ape evolution.PLoS Biol. 2004 Jul;2(7):E207. doi: 10.1371/journal.pbio.0020207. Epub 2004 Jul 13. PLoS Biol. 2004. PMID: 15252450 Free PMC article.
-
Duplication and positive selection among hominin-specific PRAME genes.BMC Genomics. 2005 Sep 13;6:120. doi: 10.1186/1471-2164-6-120. BMC Genomics. 2005. PMID: 16159394 Free PMC article.
-
Structural divergence between the human and chimpanzee genomes.Hum Genet. 2007 Feb;120(6):759-78. doi: 10.1007/s00439-006-0270-6. Epub 2006 Oct 26. Hum Genet. 2007. PMID: 17066299 Review.
-
Evolutionarily conserved low copy repeats (LCRs) in 22q11 mediate deletions, duplications, translocations, and genomic instability: an update and literature review.Genet Med. 2001 Jan-Feb;3(1):6-13. doi: 10.1097/00125817-200101000-00003. Genet Med. 2001. PMID: 11339380 Review.
Cited by
-
Transcript analysis of laser capture microdissected white matter astrocytes and higher phenol sulfotransferase 1A1 expression during autoimmune neuroinflammation.J Neuroinflammation. 2015 Jul 4;12:130. doi: 10.1186/s12974-015-0348-y. J Neuroinflammation. 2015. PMID: 26141738 Free PMC article.
-
Sulfotransferase 1A3/4 copy number variation is associated with neurodegenerative disease.Pharmacogenomics J. 2018 Apr;18(2):209-214. doi: 10.1038/tpj.2017.4. Epub 2017 Apr 4. Pharmacogenomics J. 2018. PMID: 28374858
-
Integrating protein structures and precomputed genealogies in the Magnum database: examples with cellular retinoid binding proteins.BMC Bioinformatics. 2006 Feb 23;7:89. doi: 10.1186/1471-2105-7-89. BMC Bioinformatics. 2006. PMID: 16504077 Free PMC article.
-
Sulfate activation enzymes: phylogeny and association with pyrophosphatase.J Mol Evol. 2009 Jan;68(1):1-13. doi: 10.1007/s00239-008-9181-6. Epub 2008 Dec 6. J Mol Evol. 2009. PMID: 19067028
-
Sulfotransferase gene copy number variation: pharmacogenetics and function.Cytogenet Genome Res. 2008;123(1-4):205-10. doi: 10.1159/000184710. Epub 2009 Mar 11. Cytogenet Genome Res. 2008. PMID: 19287157 Free PMC article. Review.
References
-
- Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J, Devon K, Dewar K, Doyle M, FitzHugh W, Funke R, Gage D, Harris K, Heaford A, Howland J, Kann L, Lehoczky J, LeVine R, McEwan P, McKernan K, Meldrim J, Mesirov JP, Miranda C, Morris W, Naylor J, Raymond C, Rosetti M, Santos R, Sheridan A, Sougnez C, Stange-Thomann N, Stojanovic N, Subramanian A, Wyman D, Rogers J, Sulston J, Ainscough R, Beck S, Bentley D, Burton J, Clee C, Carter N, Coulson A, Deadman R, Deloukas P, Dunham A, Dunham I, Durbin R, French L, Grafham D, Gregory S, Hubbard T, Humphray S, Hunt A, Jones M, Lloyd C, McMurray A, Matthews L, Mercer S, Milne S, Mullikin JC, Mungall A, Plumb R, Ross M, Shownkeen R, Sims S, Waterston RH, Wilson RK, Hillier LW, McPherson JD, Marra MA, Mardis ER, Fulton LA, Chinwalla AT, Pepin KH, Gish WR, Chissoe SL, Wendl MC, Delehaunty KD, Miner TL, Delehaunty A, Kramer JB, Cook LL, Fulton RS, Johnson DL, Minx PJ, Clifton SW, Hawkins T, Branscomb E, Predki P, Richardson P, Wenning S, Slezak T, Doggett N, Cheng JF, Olsen A, Lucas S, Elkin C, Uberbacher E, Frazier M, Gibbs RA, Muzny DM, Scherer SE, Bouck JB, Sodergren EJ, Worley KC, Rives CM, Gorrell JH, Metzker ML, Naylor SL, Kucherlapati RS, Nelson DL, Weinstock GM, Sakaki Y, Fujiyama A, Hattori M, Yada T, Toyoda A, Itoh T, Kawagoe C, Watanabe H, Totoki Y, Taylor T, Weissenbach J, Heilig R, Saurin W, Artiguenave F, Brottier P, Bruls T, Pelletier E, Robert C, Wincker P, Smith DR, Doucette-Stamm L, Rubenfield M, Weinstock K, Lee HM, Dubois J, Rosenthal A, Platzer M, Nyakatura G, Taudien S, Rump A, Yang H, Yu J, Wang J, Huang G, Gu J, Hood L, Rowen L, Madan A, Qin S, Davis RW, Federspiel NA, Abola AP, Proctor MJ, Myers RM, Schmutz J, Dickson M, Grimwood J, Cox DR, Olson MV, Kaul R, Shimizu N, Kawasaki K, Minoshima S, Evans GA, Athanasiou M, Schultz R, Roe BA, Chen F, Pan H, Ramser J, Lehrach H, Reinhardt R, McCombie WR, de la Bastide M, Dedhia N, Blocker H, Hornischer K, Nordsiek G, Agarwala R, Aravind L, Bailey JA, Bateman A, Batzoglou S, Birney E, Bork P, Brown DG, Burge CB, Cerutti L, Chen HC, Church D, Clamp M, Copley RR, Doerks T, Eddy SR, Eichler EE, Furey TS, Galagan J, Gilbert JG, Harmon C, Hayashizaki Y, Haussler D, Hermjakob H, Hokamp K, Jang W, Johnson LS, Jones TA, Kasif S, Kaspryzk A, Kennedy S, Kent WJ, Kitts P, Koonin EV, Korf I, Kulp D, Lancet D, Lowe TM, McLysaght A, Mikkelsen T, Moran JV, Mulder N, Pollara VJ, Ponting CP, Schuler G, Schultz J, Slater G, Smit AF, Stupka E, Szustakowski J, Thierry-Mieg D, Thierry-Mieg J, Wagner L, Wallis J, Wheeler R, Williams A, Wolf YI, Wolfe KH, Yang SP, Yeh RF, Collins F, Guyer MS, Peterson J, Felsenfeld A, Wetterstrand KA, Patrinos A, Morgan MJ, Szustakowki J, de Jong P, Catanese JJ, Osoegawa K, Shizuya H, Choi S, Chen YJ. Initial sequencing and analysis of the human genome. Nature. 2001;409:860–921. doi: 10.1038/35057062. - DOI - PubMed
-
- Cheung VG, Nowak N, Jang W, Kirsch IR, Zhao S, Chen XN, Furey TS, Kim UJ, Kuo WL, Olivier M, Conroy J, Kasprzyk A, Massa H, Yonescu R, Sait S, Thoreen C, Snijders A, Lemyre E, Bailey JA, Bruzel A, Burrill WD, Clegg SM, Collins S, Dhami P, Friedman C, Han CS, Herrick S, Lee J, Ligon AH, Lowry S, Morley M, Narasimhan S, Osoegawa K, Peng Z, Plajzer-Frick I, Quade BJ, Scott D, Sirotkin K, Thorpe AA, Gray JW, Hudson J, Pinkel D, Ried T, Rowen L, Shen-Ong GL, Strausberg RL, Birney E, Callen DF, Cheng JF, Cox DR, Doggett NA, Carter NP, Eichler EE, Haussler D, Korenberg JR, Morton CC, Albertson D, Schuler G, de Jong PJ, Trask BJ. Integration of cytogenetic landmarks into the draft sequence of the human genome. Nature. 2001;409:953–958. doi: 10.1038/35057192. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

