Subcellular localization and membrane association of SARS-CoV 3a protein
- PMID: 15763150
- PMCID: PMC7114198
- DOI: 10.1016/j.virusres.2005.01.001
Subcellular localization and membrane association of SARS-CoV 3a protein
Abstract
SARS-CoV 3a protein was a unique protein of SARS coronavirus (SARS-CoV), which was identified in SARS-CoV infected cells and SARS patients' specimen. Recent studies revealed that 3a could interact specifically with many SARS-CoV structural proteins, such as M, E and S protein. Expressed 3a protein was reported to localize to Golgi complex in SARS-CoV infected cells. In this study, it was shown that 3a protein was mainly located in Golgi apparatus with different tags at N- or C-terminus. The localization pattern was similar in different transfected cells. With the assay of truncated 3a protein, it was shown that 3a might contain three transmembrane regions, and the second or third region was properly responsible for Golgi localization. By ultra-centrifugation experiment with different extraction buffers, it was confirmed that 3a was an integral membrane protein and embedded in the phospholipid bilayer. Immunofluorescence assay indicated that 3a was co-localized with M protein in Golgi complex in co-transfected cells. These results provide a new insight for further study of the 3a protein on the pathogenesis of SARS-CoV.
Figures


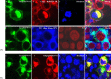





References
-
- Babia T., Ledesma M.D., Saffrich R., Kok J.W., Dotti C.G., Egea G. Endocytosis of NBD-sphingolipids in neurons: exclusion from degradative compartments and transport to the Golgi complex. Traffic. 2001;2(6):395–405. - PubMed
-
- Cole L., Davies D., Hyde G.J., Ashford A.E. ER-Tracker dye and BODIPY-brefeldin A differentiate the endoplasmic reticulum and Golgi bodies from the tubular–vacuole system in living hyphae of Pisolithus tinctorius. J. Microsc. 2000;197(3):239–249. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

