The minor capsid protein L2 contributes to two steps in the human papillomavirus type 31 life cycle
- PMID: 15767396
- PMCID: PMC1061585
- DOI: 10.1128/JVI.79.7.3938-3948.2005
The minor capsid protein L2 contributes to two steps in the human papillomavirus type 31 life cycle
Abstract
Prior studies, which have relied upon the use of pseudovirions generated in heterologous cell types, have led to sometimes conflicting conclusions regarding the role of the minor capsid protein of papillomaviruses, L2, in the viral life cycle. In this study we carry out analyses with true virus particles assembled in the natural host cell to assess L2's role in the viral infectious life cycle. For these studies we used the organotypic (raft) culture system to recapitulate the full viral life cycle of the high-risk human papillomavirus HPV31, which was either wild type or mutant for L2. After transfection, the L2 mutant HPV31 genome was able to establish itself as a nuclear plasmid in proliferating populations of poorly differentiated (basal-like) human keratinocytes and to amplify its genome to high copy number, support late viral gene expression, and cause formation of virus particles in human keratinocytes that had been induced to undergo terminal differentiation. These results indicate that aspects of both the nonproductive and productive phases of the viral life cycle occur normally in the absence of functional L2. However, upon the analysis of the virus particles generated, we found an approximate 10-fold reduction in the amount of viral DNA encapsidated into L2-deficient virions. Furthermore, there was an over-100-fold reduction in the infectivity of L2-deficient virus. Because the latter deficiency cannot be accounted for solely by the 10-fold decrease in encapsidation, we conclude that L2 contributes to at least two steps in the production of infectious virus.
Figures

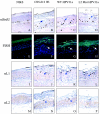




References
-
- Allen-Hoffman, B., S. Schlosser, C. Ivarie, L. Meisner, and S. O'Connor. 2000. Normal growth and differentiation in a spontaneously immortalized near-diploid human keratinocyte cell line, NIKS. J. Investig. Dermatol. 114:444-455. - PubMed
-
- Christensen, N., J. Dillner, C. Eklund, J. Carter, G. Wipf, C. Reed, N. Cladel, and D. Galloway. 1996. Surface conformational and linear epitoes on HPV-16 and HPV-18 virus like particles as defined by monoclonal antibodies. Virology 223:174-184. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

