From cyclohydrolase to oxidoreductase: discovery of nitrile reductase activity in a common fold
- PMID: 15767583
- PMCID: PMC555470
- DOI: 10.1073/pnas.0408056102
From cyclohydrolase to oxidoreductase: discovery of nitrile reductase activity in a common fold
Abstract
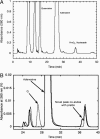
The enzyme YkvM from Bacillus subtilis was identified previously along with three other enzymes (YkvJKL) in a bioinformatics search for enzymes involved in the biosynthesis of queuosine, a 7-deazaguanine modified nucleoside found in tRNA(GUN) of Bacteria and Eukarya. Genetic analysis of ykvJKLM mutants in Acinetobacter confirmed that each was essential for queuosine biosynthesis, and the genes were renamed queCDEF. QueF exhibits significant homology to the type I GTP cyclohydrolases characterized by FolE. Given that GTP is the precursor to queuosine and that a cyclohydrolase-like reaction was postulated as the initial step in queuosine biosynthesis, QueF was proposed to be the putative cyclohydrolase-like enzyme responsible for this reaction. We have cloned the queF genes from B. subtilis and Escherichia coli and characterized the recombinant enzymes. Contrary to the predictions based on sequence analysis, we discovered that the enzymes, in fact, catalyze a mechanistically unrelated reaction, the NADPH-dependent reduction of 7-cyano-7-deazaguanineto7-aminomethyl-7-deazaguanine, a late step in the biosynthesis of queuosine. We report here in vitro and in vivo studies that demonstrate this catalytic activity, as well as preliminary biochemical and bioinformatics analysis that provide insight into the structure of this family of enzymes.
Figures

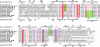


Similar articles
-
Mechanistic studies of Bacillus subtilis QueF, the nitrile oxidoreductase involved in queuosine biosynthesis.Biochemistry. 2007 Nov 6;46(44):12844-54. doi: 10.1021/bi701265r. Epub 2007 Oct 11. Biochemistry. 2007. PMID: 17929836
-
Crystallization and preliminary X-ray characterization of the nitrile reductase QueF: a queuosine-biosynthesis enzyme.Acta Crystallogr Sect F Struct Biol Cryst Commun. 2005 Oct 1;61(Pt 10):945-8. doi: 10.1107/S1744309105029246. Epub 2005 Sep 30. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2005. PMID: 16511203 Free PMC article.
-
Discovery of a new prokaryotic type I GTP cyclohydrolase family.J Biol Chem. 2006 Dec 8;281(49):37586-93. doi: 10.1074/jbc.M607114200. Epub 2006 Oct 10. J Biol Chem. 2006. PMID: 17032654
-
Biosynthesis of vitamin b2 (riboflavin).Annu Rev Nutr. 2000;20:153-67. doi: 10.1146/annurev.nutr.20.1.153. Annu Rev Nutr. 2000. PMID: 10940330 Review.
-
Genes, enzymes and coenzymes of queuosine biosynthesis in procaryotes.Biochimie. 1994;76(12):1178-82. doi: 10.1016/0300-9084(94)90047-7. Biochimie. 1994. PMID: 7748953 Review.
Cited by
-
Crystal structure of the archaeosine synthase QueF-like-Insights into amidino transfer and tRNA recognition by the tunnel fold.Proteins. 2017 Jan;85(1):103-116. doi: 10.1002/prot.25202. Epub 2016 Nov 20. Proteins. 2017. PMID: 27802572 Free PMC article.
-
7-Deazaguanine modifications protect phage DNA from host restriction systems.Nat Commun. 2019 Nov 29;10(1):5442. doi: 10.1038/s41467-019-13384-y. Nat Commun. 2019. PMID: 31784519 Free PMC article.
-
The copper-inducible cin operon encodes an unusual methionine-rich azurin-like protein and a pre-Q0 reductase in Pseudomonas putida KT2440.J Bacteriol. 2007 Jul;189(14):5361-71. doi: 10.1128/JB.00377-07. Epub 2007 May 4. J Bacteriol. 2007. PMID: 17483220 Free PMC article.
-
Radical SAM enzymes involved in the biosynthesis of purine-based natural products.Biochim Biophys Acta. 2012 Nov;1824(11):1245-53. doi: 10.1016/j.bbapap.2012.07.014. Epub 2012 Aug 3. Biochim Biophys Acta. 2012. PMID: 22902275 Free PMC article. Review.
-
Designing calcium release channel inhibitors with enhanced electron donor properties: stabilizing the closed state of ryanodine receptor type 1.Mol Pharmacol. 2012 Jan;81(1):53-62. doi: 10.1124/mol.111.074740. Epub 2011 Oct 11. Mol Pharmacol. 2012. PMID: 21989257 Free PMC article.
References
-
- Coulson, A. F. & Moult, J. (2002) Proteins 46, 61–71. - PubMed
-
- Babbitt, P. C. & Gerlt, J. A. (1997) J. Biol. Chem. 272, 30591–30594. - PubMed
-
- Chothia, C., Gough, J., Vogel, C. & Teichmann, S. A. (2003) Science 300, 1701–1703. - PubMed
-
- Teichmann, S. A., Rison, S. C., Thornton, J. M., Riley, M., Gough, J. & Chothia, C. (2001) J. Mol. Biol. 311, 693–708. - PubMed
-
- Copley, R. R. & Bork, P. (2000) J. Mol. Biol. 303, 627–641. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

