Class II histone deacetylases: from sequence to function, regulation, and clinical implication
- PMID: 15798178
- PMCID: PMC1069616
- DOI: 10.1128/MCB.25.8.2873-2884.2005
Class II histone deacetylases: from sequence to function, regulation, and clinical implication
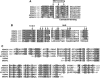
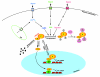
Figures

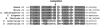
Similar articles
-
Deacetylase enzymes: biological functions and the use of small-molecule inhibitors.Chem Biol. 2002 Jan;9(1):3-16. doi: 10.1016/s1074-5521(02)00092-3. Chem Biol. 2002. PMID: 11841934 Review. No abstract available.
-
Identification of HDAC10, a novel class II human histone deacetylase containing a leucine-rich domain.Nucleic Acids Res. 2002 Mar 1;30(5):1114-23. doi: 10.1093/nar/30.5.1114. Nucleic Acids Res. 2002. PMID: 11861901 Free PMC article.
-
Histone deacetylase inhibitors and demethylating agents: clinical development of histone deacetylase inhibitors for cancer therapy.Cancer J. 2007 Jan-Feb;13(1):30-9. doi: 10.1097/PPO.0b013e31803c73cc. Cancer J. 2007. PMID: 17464244 Review.
-
Class IIa histone deacetylases: regulating the regulators.Oncogene. 2007 Aug 13;26(37):5450-67. doi: 10.1038/sj.onc.1210613. Oncogene. 2007. PMID: 17694086 Review.
-
HDAC8: a multifaceted target for therapeutic interventions.Trends Pharmacol Sci. 2015 Jul;36(7):481-92. doi: 10.1016/j.tips.2015.04.013. Epub 2015 May 23. Trends Pharmacol Sci. 2015. PMID: 26013035 Review.
Cited by
-
Transcriptional pathways and potential therapeutic targets in the regulation of Ncx1 expression in cardiac hypertrophy and failure.Adv Exp Med Biol. 2013;961:125-35. doi: 10.1007/978-1-4614-4756-6_11. Adv Exp Med Biol. 2013. PMID: 23224875 Free PMC article. Review.
-
Role of histone deacetylase inhibitors in non-neoplastic diseases.Heliyon. 2024 Jul 2;10(13):e33997. doi: 10.1016/j.heliyon.2024.e33997. eCollection 2024 Jul 15. Heliyon. 2024. PMID: 39071622 Free PMC article. Review.
-
Histone Deacetylases Exert Class-Specific Roles in Conditioning the Brain and Heart Against Acute Ischemic Injury.Front Neurol. 2015 Jun 30;6:145. doi: 10.3389/fneur.2015.00145. eCollection 2015. Front Neurol. 2015. PMID: 26175715 Free PMC article. Review.
-
Histone deacetylase inhibitors synergize p300 autoacetylation that regulates its transactivation activity and complex formation.Cancer Res. 2007 Mar 1;67(5):2256-2264. doi: 10.1158/0008-5472.CAN-06-3985. Cancer Res. 2007. PMID: 17332356 Free PMC article.
-
Effects of histone deacetylase inhibitors on HIF-1.Cell Cycle. 2006 Nov 1;5(21):2430-5. doi: 10.4161/cc.5.21.3409. Epub 2006 Sep 13. Cell Cycle. 2006. PMID: 17102633 Free PMC article. Review.
References
-
- Amann, J. M., J. Nip, D. K. Strom, B. Lutterbach, H. Harada, N. Lenny, J. R. Downing, S. Meyers, and S. W. Hiebert. 2001. ETO, a target of t(8;21) in acute leukemia, makes distinct contacts with multiple histone deacetylases and binds mSin3A through its oligomerization domain. Mol. Cell. Biol. 21:6470-6483. - PMC - PubMed
-
- Amerik, A. Y., S. J. Li, and M. Hochstrasser. 2000. Analysis of the deubiquitinating enzymes of the yeast Saccharomyces cerevisiae. Biol. Chem. 381:981-992. - PubMed
-
- Bakin, R. E., and M. O. Jung. 2004. Cytoplasmic sequestration of HDAC7 from mitochondrial and nuclear compartments upon initiation of apoptosis. J. Biol. Chem. 279:51218-51225. - PubMed
-
- Barlow, A. L., C. M. van Drunen, C. A. Johnson, S. Tweedie, A. Bird, and B. M. Turner. 2001. dSIR2 and dHDAC6: two novel, inhibitor-resistant deacetylases in Drosophila melanogaster. Exp. Cell Res. 265:90-103. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
