A novel SR-related protein is required for the second step of Pre-mRNA splicing
- PMID: 15798186
- PMCID: PMC1069619
- DOI: 10.1128/MCB.25.8.2969-2980.2005
A novel SR-related protein is required for the second step of Pre-mRNA splicing
Abstract
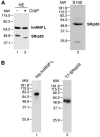
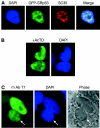
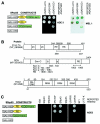
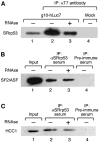
The SR family proteins and SR-related polypeptides are important regulators of pre-mRNA splicing. A novel SR-related protein of an apparent molecular mass of 53 kDa was isolated in a gene trap screen that identifies proteins which localize to the nuclear speckles. This novel protein possesses an arginine- and serine-rich domain and was termed SRrp53 (for SR-related protein of 53 kDa). In support for a role of this novel RS-containing protein in pre-mRNA splicing, we identified the mouse ortholog of the Saccharomyces cerevisiae U1 snRNP-specific protein Luc7p and the U2AF65-related factor HCC1 as interacting proteins. In addition, SRrp53 is able to interact with some members of the SR family of proteins and with U2AF35 in a yeast two-hybrid system and in cell extracts. We show that in HeLa nuclear extracts immunodepleted of SRrp53, the second step of pre-mRNA splicing is blocked, and recombinant SRrp53 is able to restore splicing activity. SRrp53 also regulates alternative splicing in a concentration-dependent manner. Taken together, these results suggest that SRrp53 is a novel SR-related protein that has a role both in constitutive and in alternative splicing.
Figures


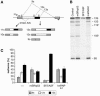
References
-
- Apweiler, R., T. K. Attwood, A. Bairoch, A. Bateman, E. Birney, M. Biswas, P. Bucher, L. Cerutti, F. Corpet, M. D. Croning, R. Durbin, L. Falquet, W. Fleischmann, J. Gouzy, H. Hermjakob, N. Hulo, I. Jonassen, D. Kahn, A. Kanapin, Y. Karavidopoulou, R. Lopez, B. Marx, N. J. Mulder, T. M. Oinn, M. Pagni, F. Servant, C. J. Sigrist, and E. M. Zdobnov for the InterPro Consortium. 2000. InterPro—an integrated documentation resource for protein families, domains and functional sites. Bioinformatics 16:1145-1150. - PubMed
-
- Blencowe, B. J. 2000. Exonic splicing enhancers: mechanism of action, diversity and role in human genetic diseases. Trends Biochem. Sci. 25:106-110. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
