Topology of transmembrane segments 1-4 in the human chloride/bicarbonate anion exchanger 1 (AE1) by scanning N-glycosylation mutagenesis
- PMID: 15804238
- PMCID: PMC1184569
- DOI: 10.1042/BJ20050315
Topology of transmembrane segments 1-4 in the human chloride/bicarbonate anion exchanger 1 (AE1) by scanning N-glycosylation mutagenesis
Abstract
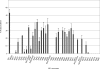
Human AE1 (anion exchanger 1), or Band 3, is an abundant membrane glycoprotein found in the plasma membrane of erythrocytes. The physiological role of the protein is to carry out chloride/bicarbonate exchange across the plasma membrane, a process that increases the carbon-dioxide-carrying capacity of blood. To study the topology of TMs (transmembrane segments) 1-4, a series of scanning N-glycosylation mutants were created spanning the region from EC (extracellular loop) 1 to EC2 in full-length AE1. These constructs were expressed in HEK-293 (human embryonic kidney) cells, and their N-glycosylation efficiencies were determined. Unexpectedly, positions within putative TMs 2 and 3 could be efficiently glycosylated. In contrast, the same positions were very poorly glycosylated when present in mutant AE1 with the SAO (Southeast Asian ovalocytosis) deletion (DeltaA400-A408) in TM1. These results suggest that the TM2-3 region of AE1 may become transiently exposed to the endoplasmic reticulum lumen during biosynthesis, and that there is a competition between proper folding of the region into the membrane and N-glycosylation at introduced sites. The SAO deletion disrupts the proper integration of TMs 1-2, probably leaving the region exposed to the cytosol. As a result, engineered N-glycosylation acceptor sites in TM2-3 could not be utilized by the oligosaccharyltransferase in this mutant form of AE1. The properties of TM2-3 suggest that these segments form a re-entrant loop in human AE1.
Figures


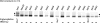

Similar articles
-
Transmembrane folding of the human erythrocyte anion exchanger (AE1, Band 3) determined by scanning and insertional N-glycosylation mutagenesis.Biochem J. 1999 Apr 15;339 ( Pt 2)(Pt 2):269-79. Biochem J. 1999. PMID: 10191257 Free PMC article.
-
Membrane integration and topology of the first transmembrane segment in normal and Southeast Asian ovalocytosis human erythrocyte anion exchanger 1.Mol Membr Biol. 2005 May-Jun;22(3):203-14. doi: 10.1080/09687860500093115. Mol Membr Biol. 2005. PMID: 16096263
-
Palmitoylation is not required for trafficking of human anion exchanger 1 to the cell surface.Biochem J. 2004 Mar 15;378(Pt 3):1015-21. doi: 10.1042/BJ20030847. Biochem J. 2004. PMID: 14640982 Free PMC article.
-
Structure of the band 3 transmembrane domain.Cell Mol Biol (Noisy-le-grand). 1996 Nov;42(7):1041-51. Cell Mol Biol (Noisy-le-grand). 1996. PMID: 8960779 Review.
-
Band 3, the human red cell chloride/bicarbonate anion exchanger (AE1, SLC4A1), in a structural context.Biochim Biophys Acta. 2016 Jul;1858(7 Pt A):1507-32. doi: 10.1016/j.bbamem.2016.03.030. Epub 2016 Apr 6. Biochim Biophys Acta. 2016. PMID: 27058983 Review.
Cited by
-
Topology of NBCe1 protein transmembrane segment 1 and structural effect of proximal renal tubular acidosis (pRTA) S427L mutation.J Biol Chem. 2013 Mar 15;288(11):7894-7906. doi: 10.1074/jbc.M112.404533. Epub 2013 Jan 28. J Biol Chem. 2013. PMID: 23362273 Free PMC article.
-
The divergence, actions, roles, and relatives of sodium-coupled bicarbonate transporters.Physiol Rev. 2013 Apr;93(2):803-959. doi: 10.1152/physrev.00023.2012. Physiol Rev. 2013. PMID: 23589833 Free PMC article. Review.
-
NBCe1-A Transmembrane Segment 1 Lines the Ion Translocation Pathway.J Biol Chem. 2009 Mar 27;284(13):8918-29. doi: 10.1074/jbc.M806674200. Epub 2009 Jan 21. J Biol Chem. 2009. PMID: 19158093 Free PMC article.
-
Mutation of N-linked glycosylation at Asn548 in CD133 decreases its ability to promote hepatoma cell growth.Oncotarget. 2015 Aug 21;6(24):20650-60. doi: 10.18632/oncotarget.4115. Oncotarget. 2015. PMID: 26029999 Free PMC article.
-
Effect of the Southeast Asian Ovalocytosis Deletion on the Conformational Dynamics of Signal-Anchor Transmembrane Segment 1 of Red Cell Anion Exchanger 1 (AE1, Band 3, or SLC4A1).Biochemistry. 2017 Feb 7;56(5):712-722. doi: 10.1021/acs.biochem.6b00966. Epub 2017 Jan 23. Biochemistry. 2017. PMID: 28068080 Free PMC article.
References
-
- Grinstein S., Ship S., Rothstein A. Anion transport in relation to proteolytic dissection of band 3 protein. Biochim. Biophys. Acta. 1978;507:294–304. - PubMed
-
- Low P. S. Structure and function of the cytoplasmic domain of band 3: center of erythrocyte membrane–peripheral protein interactions. Biochim. Biophys. Acta. 1986;864:145–167. - PubMed
-
- Tanner M. J. A. Band 3 anion exchanger and its involvement in erythrocyte and kidney disorders. Curr. Opin. Hematol. 2002;9:133–139. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

