Translational control of tetracycline resistance and conjugation in the Bacteroides conjugative transposon CTnDOT
- PMID: 15805513
- PMCID: PMC1070362
- DOI: 10.1128/JB.187.8.2673-2680.2005
Translational control of tetracycline resistance and conjugation in the Bacteroides conjugative transposon CTnDOT
Abstract
The tetQ-rteA-rteB operon of the Bacteroides conjugative transposon CTnDOT is responsible for tetracycline control of the excision and transfer of CTnDOT. Previous studies revealed that tetracycline control of this operon occurred at the translational level and involved a hairpin structure located within the 130-base leader sequence that lies between the promoter of tetQ and the start codon of the gene. This hairpin structure is formed by two sequences, designated Hp1 and Hp8. Hp8 contains the ribosome binding site for tetQ. Examination of the leader region sequence revealed three sequences that might encode a leader peptide. One was only 3 amino acids long. The other two were 16 amino acids long. By introducing stop codons into the peptide coding regions, we have now shown that the 3-amino-acid peptide is the one that is essential for tetracycline control. Between Hp1 and Hp8 lies an 85-bp region that contains other possible RNA hairpin structures. Deletion analysis of this intervening DNA segment has now identified a sequence, designated Hp2, which is essential for tetracycline regulation. This sequence could form a short hairpin structure with Hp1. Mutations that made the Hp1-Hp2 structure more stable caused nearly constitutively high expression of the operon. Thus, stalling of ribosomes on the 3-amino-acid leader peptide could favor formation of the Hp1-Hp2 structure and thus preclude formation of the Hp1-Hp8 structure, releasing the ribosome binding site of tetQ. Finally, comparison of the CTnDOT tetQ leader regions with upstream regions of five tetQ genes found in other elements reveals that the sequences are virtually identical, suggesting that translational attenuation is responsible for control of tetracycline resistance in these other cases as well.
Figures
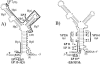
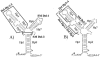
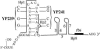

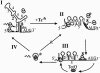
Similar articles
-
Regulation of a Bacteroides operon that controls excision and transfer of the conjugative transposon CTnDOT.J Bacteriol. 2004 May;186(9):2548-57. doi: 10.1128/JB.186.9.2548-2557.2004. J Bacteriol. 2004. PMID: 15090494 Free PMC article.
-
Regulation of excision genes of the Bacteroides conjugative transposon CTnDOT.J Bacteriol. 2005 Aug;187(16):5732-41. doi: 10.1128/JB.187.16.5732-5741.2005. J Bacteriol. 2005. PMID: 16077120 Free PMC article.
-
An unexpected effect of tetracycline concentration: growth phase-associated excision of the Bacteroides mobilizable transposon NBU1.J Bacteriol. 2009 Feb;191(3):1078-82. doi: 10.1128/JB.00637-08. Epub 2008 Oct 24. J Bacteriol. 2009. PMID: 18952794 Free PMC article.
-
Regulation of CTnDOT conjugative transfer is a complex and highly coordinated series of events.mBio. 2013 Oct 29;4(6):e00569-13. doi: 10.1128/mBio.00569-13. mBio. 2013. PMID: 24169574 Free PMC article. Review.
-
Chromosomal gene transfer elements of the Bacteroides group.Eur J Clin Microbiol Infect Dis. 1992 Nov;11(11):1032-8. doi: 10.1007/BF01967795. Eur J Clin Microbiol Infect Dis. 1992. PMID: 1338314 Review.
Cited by
-
Microbiological effects of sublethal levels of antibiotics.Nat Rev Microbiol. 2014 Jul;12(7):465-78. doi: 10.1038/nrmicro3270. Epub 2014 May 27. Nat Rev Microbiol. 2014. PMID: 24861036 Review.
-
The frequency of tetracycline resistance genes co-detected with respiratory pathogens: a database mining study uncovering descriptive trends throughout the United States.BMC Infect Dis. 2014 Aug 25;14:460. doi: 10.1186/1471-2334-14-460. BMC Infect Dis. 2014. PMID: 25152108 Free PMC article.
-
Development of an IPTG inducible expression vector adapted for Bacteroides fragilis.Plasmid. 2012 Sep;68(2):86-92. doi: 10.1016/j.plasmid.2012.03.002. Epub 2012 Apr 1. Plasmid. 2012. PMID: 22487080 Free PMC article.
-
Antibiotics Promote Escherichia coli-Pseudomonas aeruginosa Conjugation through Inhibiting Quorum Sensing.Antimicrob Agents Chemother. 2017 Nov 22;61(12):e01284-17. doi: 10.1128/AAC.01284-17. Print 2017 Dec. Antimicrob Agents Chemother. 2017. PMID: 28993333 Free PMC article.
-
The Xis2d protein of CTnDOT binds to the intergenic region between the mob and tra operons.Plasmid. 2015 Sep;81:63-71. doi: 10.1016/j.plasmid.2015.07.002. Epub 2015 Jul 23. Plasmid. 2015. PMID: 26212728 Free PMC article.
References
-
- Bayley, D. P., E. R. Rocha, and C. J. Smith. 2000. Analysis of cepA and other Bacteroides fragilis genes reveals a unique promoter structure. FEMS Microbiol. Lett. 193:149-154. - PubMed
-
- Berens, C., and W. Hillen. 2003. Gene regulation by tetracyclines. Constraints of resistance regulation in bacteria shape TetR for application in eukaryotes. Eur. J. Biochem. 270:3109-3121. - PubMed
-
- Boyer, H. B., and D. Roulland-Dussoix. 1969. A complementation analysis of the restriction and modification system of DNA in Escherichia coli. J. Mol. Biol. 41:459-472. - PubMed
-
- Cheng, Q., Y. Sutanto, N. B. Shoemaker, J. F. Gardner, and A. A. Salyers. 2001. Identification of genes required for the excision of CTnDOT, a Bacteroides conjugative transposon. Mol. Microbiol. 41:625-632. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

