Diversity and phylogenetic affiliations of morphologically conspicuous large filamentous bacteria occurring in the pelagic zones of a broad spectrum of freshwater habitats
- PMID: 15812022
- PMCID: PMC1082555
- DOI: 10.1128/AEM.71.4.1931-1940.2005
Diversity and phylogenetic affiliations of morphologically conspicuous large filamentous bacteria occurring in the pelagic zones of a broad spectrum of freshwater habitats
Abstract
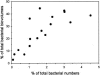
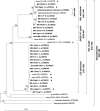
Filamentous bacteria with a conspicuous morphology were found in the majority of the bacterioplankton samples from a variety of freshwater habitats that were studied. These heterotrophic filaments typically account for < 1 to 11% of the total number of bacteria. The biovolume of this morphotype can exceed 40% of the biovolume for all bacteria. Surprisingly, we found hardly any data on these morphologically conspicuous filaments in the literature. Mixed cultures containing these filamentous bacteria were established by cultivation and isolation experiments with samples from different freshwater lakes. Nearly full-length 16S rRNA gene sequences were obtained from several mixed cultures and environmental samples from habitats in Europe, Africa, China, Australia, and New Zealand. Phylogenetic analysis of the sequences showed that three groups form a single monophyletic cluster, the SOL cluster, in the family Saprospiraceae. We developed a set of six nested probes for fluorescence in situ hybridization. Of the six probes, one probe was specific for Haliscomenobacter hydrossis, three probes were specific for the three subclusters (each probe was specific for one subcluster), one probe was specific for the entire SOL cluster, and another probe targeted almost the entire Saprospiraceae family. Specific hybridization of environmental samples and enrichments showed that the members of the three subclusters exhibited the same filamentous morphology. So far, using the subcluster-specific probes, we have not been able to detect any bacteria with a differing morphology. We conclude that the SOL cluster bacteria are an integral part of bacterioplankton in many freshwater habitats. They potentially account for a large fraction of the total bacterial biomass but have been underrepresented in molecular diversity studies so far.
Figures


Similar articles
-
A guide to the natural history of freshwater lake bacteria.Microbiol Mol Biol Rev. 2011 Mar;75(1):14-49. doi: 10.1128/MMBR.00028-10. Microbiol Mol Biol Rev. 2011. PMID: 21372319 Free PMC article. Review.
-
Ecological differentiation within a cosmopolitan group of planktonic freshwater bacteria (SOL cluster, Saprospiraceae, Bacteroidetes).Appl Environ Microbiol. 2005 Oct;71(10):5900-7. doi: 10.1128/AEM.71.10.5900-5907.2005. Appl Environ Microbiol. 2005. PMID: 16204503 Free PMC article.
-
'Candidatus Aquirestis calciphila' and 'Candidatus Haliscomenobacter calcifugiens', filamentous, planktonic bacteria inhabiting natural lakes.Int J Syst Evol Microbiol. 2007 May;57(Pt 5):936-940. doi: 10.1099/ijs.0.64807-0. Int J Syst Evol Microbiol. 2007. PMID: 17473236
-
Comparative 16S rRNA analysis of lake bacterioplankton reveals globally distributed phylogenetic clusters including an abundant group of actinobacteria.Appl Environ Microbiol. 2000 Nov;66(11):5053-65. doi: 10.1128/AEM.66.11.5053-5065.2000. Appl Environ Microbiol. 2000. PMID: 11055963 Free PMC article.
-
Identification and characterization of ecologically significant prokaryotes in the sediment of freshwater lakes: molecular and cultivation studies.FEMS Microbiol Rev. 2000 Dec;24(5):573-90. doi: 10.1111/j.1574-6976.2000.tb00559.x. FEMS Microbiol Rev. 2000. PMID: 11077151 Review.
Cited by
-
The balance between deterministic and stochastic processes in structuring lake bacterioplankton community over time.Mol Ecol. 2020 Aug;29(16):3117-3130. doi: 10.1111/mec.15538. Epub 2020 Jul 24. Mol Ecol. 2020. PMID: 32628343 Free PMC article.
-
Diversity and Ecophysiology of the Genus OLB8 and Other Abundant Uncultured Saprospiraceae Genera in Global Wastewater Treatment Systems.Front Microbiol. 2022 Jul 8;13:917553. doi: 10.3389/fmicb.2022.917553. eCollection 2022. Front Microbiol. 2022. PMID: 35875537 Free PMC article.
-
A guide to the natural history of freshwater lake bacteria.Microbiol Mol Biol Rev. 2011 Mar;75(1):14-49. doi: 10.1128/MMBR.00028-10. Microbiol Mol Biol Rev. 2011. PMID: 21372319 Free PMC article. Review.
-
Recurrent seasonal variations in abundance and composition of filamentous SOL cluster bacteria (Saprospiraceae, Bacteroidetes) in oligomesotrophic Lake Mondsee (Austria).Appl Environ Microbiol. 2006 Jul;72(7):4704-12. doi: 10.1128/AEM.02935-05. Appl Environ Microbiol. 2006. PMID: 16820462 Free PMC article.
-
Identification and ecophysiological characterization of epiphytic protein-hydrolyzing saprospiraceae ("Candidatus Epiflobacter" spp.) in activated sludge.Appl Environ Microbiol. 2008 Apr;74(7):2229-38. doi: 10.1128/AEM.02502-07. Epub 2008 Feb 8. Appl Environ Microbiol. 2008. PMID: 18263744 Free PMC article.
References
-
- Andersson, A., U. Larsson, and A. Hagström. 1986. Size-selective grazing by a microflagellate on pelagic bacteria. Mar. Ecol. Prog. Ser. 33:51-57.
-
- Andreatta, S. 2001. Cytometry of aquatic bacteria: analyses at the community and subgroup level. Ph.D. thesis. University of Innsbruck, Innsbruck, Austria.
-
- Bahr, M., J. E. Hobbie, and M. L. Sogin. 1996. Bacterial diversity in an arctic lake: a freshwater SAR11 cluster. Aquat. Microb. Ecol. 11:271-277.
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

