Tandem oligonucleotide synthesis using linker phosphoramidites
- PMID: 15814811
- PMCID: PMC1074720
- DOI: 10.1093/nar/gki333
Tandem oligonucleotide synthesis using linker phosphoramidites
Abstract
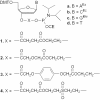
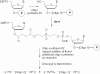
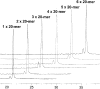
Multiple oligonucleotides of the same or different sequence, linked end-to-end in tandem can be synthesized in a single automated synthesis. A linker phosphoramidite [R. T. Pon and S. Yu (2004) Nucleic Acids Res., 32, 623-631] is added to the 5'-terminal OH end of a support-bound oligonucleotide to introduce a cleavable linkage (succinic acid plus sulfonyldiethanol) and the 3'-terminal base of the new sequence. Conventional phosphoramidites are then used for the rest of the sequence. After synthesis, treatment with ammonium hydroxide releases the oligonucleotides from the support and cleaves the linkages between each sequence. Mixtures of one oligonucleotide with both 5'- and 3'-terminal OH ends and other oligonucleotides with 5'-phosphorylated and 3'-OH ends are produced, which are deprotected and worked up as a single product. Tandem synthesis can be used to make pairs of PCR primers, sets of cooperative oligonucleotides or multiple copies of the same sequence. When tandem synthesis is used to make two self-complementary sequences, double-stranded structures spontaneously form after deprotection. Tandem synthesis of oligonucleotide chains containing up to six consecutive 20mer (120 bases total), various trinucleotide codons and primer pairs for PCR, or self-complementary strands for in situ formation of double-stranded DNA fragments has been demonstrated.
Figures
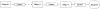



References
-
- Pon R.T., Yu S., Sanghvi Y.S. Multiple oligonucleotide synthesis in tandem on solid-phase supports for small and large scale synthesis. Nucleosides Nucleotides Nucleic Acids. 2001;20:985–989. - PubMed
-
- Pon R.T., Yu S., Sanghvi Y.S. Tandem oligonucleotide synthesis on solid-phase supports for the production of multiple oligonucleotides. J. Org. Chem. 2002;67:856–864. - PubMed
-
- Yost C.K., Nattress F.M. The use of multiplex PCR reactions to characterize populations of lactic acid bacteria associated with meat spoilage. [Erratum (2001) Lett. Appl. Microbiol., 32, 368.] Lett. Appl. Microbiol. 2000;31:129–133. - PubMed
-
- Makridakis N.M., Reichardt K.V. Multiplex automated primer extension analysis: simultaneous genotyping of several polymorphisms. Biotechniques. 2001;31:1374–1380. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

