Dynamic range of hepatitis C virus RNA quantification with the Cobas Ampliprep-Cobas Amplicor HCV Monitor v2.0 assay
- PMID: 15814982
- PMCID: PMC1081339
- DOI: 10.1128/JCM.43.4.1669-1673.2005
Dynamic range of hepatitis C virus RNA quantification with the Cobas Ampliprep-Cobas Amplicor HCV Monitor v2.0 assay
Abstract
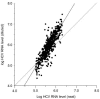
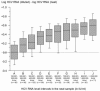
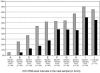
Accurate quantification of hepatitis C virus (HCV) RNA is needed in clinical practice to decide whether to continue or stop pegylated interferon-alpha-ribavirin combination therapy at week 12 of treatment for patients with chronic hepatitis C. Currently the HCV RNA quantification assay most widely used worldwide is the Amplicor HCV Monitor v2.0 assay (Roche Molecular Systems, Pleasanton, Calif.). The HCV RNA extraction step can be automated in the Cobas Ampliprep device. In this work, we show that the dynamic range of HCV RNA quantification of the Cobas Ampliprep/Cobas Amplicor HCV Monitor v2.0 procedure is 600 to 200,000 HCV RNA IU/ml (2.8 to 5.3 log IU/ml), which does not cover the full range of HCV RNA levels in infected patients. Any sample containing more than 200,000 IU/ml (5.3 log IU/ml) must thus be retested after dilution for accurate quantification. These results emphasize the need for commercial HCV RNA quantification assays with a broader range of linear quantification, such as real-time PCR-based assays.
Figures
Similar articles
-
Comparison of qualitative (COBAS AMPLICOR HCV 2.0 versus VERSANT HCV RNA) and quantitative (COBAS AMPLICOR HCV monitor 2.0 versus VERSANT HCV RNA 3.0) assays for hepatitis C virus (HCV) RNA detection and quantification: impact on diagnosis and treatment of HCV infections.J Clin Microbiol. 2005 Jun;43(6):2590-7. doi: 10.1128/JCM.43.6.2590-2597.2005. J Clin Microbiol. 2005. PMID: 15956369 Free PMC article.
-
Assessment of early virological response to antiviral therapy by comparing four assays for HCV RNA quantitation using the international unit standard: implications for clinical management of patients with chronic hepatitis C virus infection.J Med Virol. 2006 Feb;78(2):208-15. doi: 10.1002/jmv.20529. J Med Virol. 2006. PMID: 16372298 Clinical Trial.
-
Evaluation of the clinical usefulness of COBAS AMPLICOR HCV MONITOR assay (ver2.0): Comparison with AMPLICOR HCV MONITOR assay (ver1.0) and HCV core protein level.J Med Virol. 2002 Nov;68(3):343-51. doi: 10.1002/jmv.10209. J Med Virol. 2002. PMID: 12226820
-
Automation of laboratory testing for infectious diseases using the polymerase chain reaction-- our past, our present, our future.J Clin Virol. 2001 Jan;20(1-2):1-6. doi: 10.1016/s1386-6532(00)00148-7. J Clin Virol. 2001. PMID: 11163576 Review.
-
Profile of Roche's cobas® HCV tests.Expert Rev Mol Diagn. 2017 Apr;17(4):311-319. doi: 10.1080/14737159.2017.1293526. Epub 2017 Feb 22. Expert Rev Mol Diagn. 2017. PMID: 28277143 Review.
Cited by
-
Sample-to-Answer Droplet Magnetofluidic Platform for Point-of-Care Hepatitis C Viral Load Quantitation.Sci Rep. 2018 Jun 28;8(1):9793. doi: 10.1038/s41598-018-28124-3. Sci Rep. 2018. PMID: 29955160 Free PMC article.
-
Seropositive for hepatitis B and C viruses is associated with the risk of decreased bone mineral density in adults: An analysis of studies from the NHANES database.Front Med (Lausanne). 2023 Mar 22;10:1120083. doi: 10.3389/fmed.2023.1120083. eCollection 2023. Front Med (Lausanne). 2023. PMID: 37035336 Free PMC article.
-
Second-generation Cobas AmpliPrep/Cobas TaqMan HCV quantitative test for viral load monitoring: a novel dual-probe assay design.J Clin Microbiol. 2013 Feb;51(2):571-7. doi: 10.1128/JCM.01784-12. Epub 2012 Dec 12. J Clin Microbiol. 2013. PMID: 23241371 Free PMC article.
-
Real-time PCR assays for hepatitis C virus (HCV) RNA quantitation are adequate for clinical management of patients with chronic HCV infection.J Clin Microbiol. 2006 Jul;44(7):2507-11. doi: 10.1128/JCM.00163-06. J Clin Microbiol. 2006. PMID: 16825372 Free PMC article.
-
Broad differences between the COBAS ampliprep total nucleic acid isolation-COBAS TaqMan 48 hepatitis C virus (HCV) and COBAS HCV monitor v2.0 assays for quantification of serum HCV RNA of non-1 genotypes.J Clin Microbiol. 2006 Apr;44(4):1602-3. doi: 10.1128/JCM.44.4.1602-1603.2006. J Clin Microbiol. 2006. PMID: 16597910 Free PMC article. No abstract available.
References
-
- Consensus panel. 2002. National Institutes of Health Consensus Development Conference Statement: Management of hepatitis C: 2002—June 10-12, 2002. Hepatology 36:S3-S20. - PubMed
-
- Enomoto, M., S. Nishiguchi, S. Shiomi, M. Tanaka, K. Fukuda, T. Ueda, A. Tamori, D. Habu, T. Takeda, Y. Yano, and S. Otani. 2001. Comparison of real-time quantitative polymerase chain reaction with three other assays for quantitation of hepatitis C virus. J. Gastroenterol. Hepatol. 16:904-909. - PubMed
-
- Lee, S. C., A. Antony, N. Lee, J. Leibow, J. Q. Yang, S. Soviero, K. Gutekunst, and M. Rosenstraus. 2000. Improved version 2.0 qualitative and quantitative Amplicor reverse transcription-PCR tests for hepatitis C virus RNA: calibration to international units, enhanced genotype reactivity, and performance characteristics. J. Clin. Microbiol. 38:4171-4179. - PMC - PubMed
-
- Neumann, A. U., E. Hagai, S. W. Schalm, M. von Wagner, G. Germanidis, Y. Lurie, G. Missale, M. Martell, J. M. Vrolijk, C. Hezode, G. Norkrans, F. Negro, A. Soulier, E. Verheij-Hart, G. Colucci, C. Ferrari, S. Zeuzem, and J. M. Pawlotsky. 2003. Early viral kinetics prediction of sustained virological response after 1 or 4 weeks of peginterferon alfa-2a and ribavirin therapy (DITTO project). Hepatology 38:248A.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

