NMR chemical shift and relaxation measurements provide evidence for the coupled folding and binding of the p53 transactivation domain
- PMID: 15824059
- PMCID: PMC1075921
- DOI: 10.1093/nar/gki336
NMR chemical shift and relaxation measurements provide evidence for the coupled folding and binding of the p53 transactivation domain
Abstract
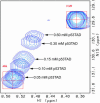
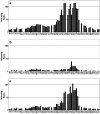
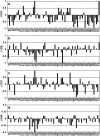
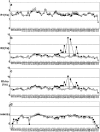
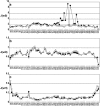
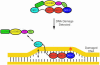
The interaction between the acidic transactivation domain of the human tumor suppressor protein p53 (p53TAD) and the 70 kDa subunit of human replication protein A (hRPA70) was investigated using heteronuclear magnetic resonance spectroscopy. A 1H-15N heteronuclear single quantum coherence (HSQC) titration experiment was performed on a 15N-labeled fragment of hRPA70, containing the N-terminal 168 residues (hRPA701-168) and p53TAD. HRPA701-168 residues important for binding were identified and found to be localized to a prominent basic cleft. This binding site overlapped with a previously identified single-stranded DNA-binding site, suggesting that a competitive binding mechanism may regulate the formation of p53TAD-hRPA70 complex. The amide 1H and 15N chemical shifts of an uniformly 15N-labeled sample of p53TAD were also monitored before and after the addition of unlabeled hRPA701-168. In the presence of unlabeled hRPA701-168, resonance lineshapes increased and corresponding intensity reductions were observed for specific p53TAD residues. The largest intensity reductions were observed for p53TAD residues 42-56. Minimal binding was observed between p53TAD and a mutant form of hRPA701-168, where the basic cleft residue R41 was changed to a glutamic acid (R41E), demonstrating that ionic interactions play an important role in specifying the binding interface. The region of p53TAD most affected by binding hRPA701-168 was found to have some residual alpha helical and beta strand structure; however, this structure was not stabilized by binding hRPA701-168. 15N relaxation experiments were performed to monitor changes in backbone dynamics of p53TAD when bound to hRPA701-168. Large changes in both the transverse (R2) and rotating frame (R1) relaxation rates were observed for a subset of the p53TAD residues that had 1H-15N HSQC resonance intensity reductions during the complex formation. The folding of p53TAD upon complex formation is suggested by the pattern of changes observed for both R2 and R1. A model that couples the formation of a weak encounter complex between p53TAD and hRPA701-168 to the folding of p53TAD is discussed in the context of a functional role for the p53-hRPA70 complex in DNA repair.
Figures



References
-
- Friedberg E.C. Correcting the Blueprint of Life: An Historical Account of the Discovery of DNA Repair Mechanisms. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press; 1997. pp. 17–19. 91–96.
-
- Friedberg E.C., Walker G.C., Siede W. DNA Repair and Mutagenesis. Washington, DC: ASM Press; 1995. pp. 1–29. 1–92.
-
- Lindahl T., Karran P., Wood R.D. DNA excision repair pathways. Curr. Opin. Genet. Dev. 1997;7:158–169. - PubMed
-
- Wallace S.S., Van Houten B., Kow Y.W. DNA Damage: Effect on DNA Structure and Protein Recognition. Annals of the New York Academy of Sciences; 1994. pp. 92–105.
-
- Wold M.S. Replication protein A: a heterotrimeric, single-stranded DNA-binding protein required for eukaryotic DNA metabolism. Annu. Rev. Biochem. 1997;66:61–92. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

