Structure and assembly of a T=1 virus-like particle in BK polyomavirus
- PMID: 15827148
- PMCID: PMC1082729
- DOI: 10.1128/JVI.79.9.5337-5345.2005
Structure and assembly of a T=1 virus-like particle in BK polyomavirus
Abstract
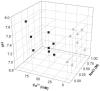
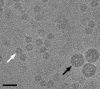
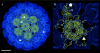
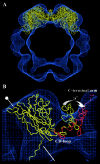
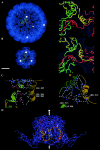
In polyomaviruses the pentameric capsomers are interlinked by the long C-terminal arm of the structural protein VP1. The T=7 icosahedral structure of these viruses is possible due to an intriguing adaptability of this linker arm to the different local environments in the capsid. To explore the assembly process, we have compared the structure of two virus-like particles (VLPs) formed, as we found, in a calcium-dependent manner by the VP1 protein of human polyomavirus BK. The structures were determined using electron cryomicroscopy (cryo-EM), and the three-dimensional reconstructions were interpreted by atomic modeling. In the small VP1 particle, 26.4 nm in diameter, the pentameric capsomers form an icosahedral T=1 surface lattice with meeting densities at the threefold axes that interlinked three capsomers. In the larger particle, 50.6 nm in diameter, the capsomers form a T=7 icosahedral shell with three unique contacts. A folding model of the BKV VP1 protein was obtained by alignment with the VP1 protein of simian virus 40 (SV40). The model fitted well into the cryo-EM density of the T=7 particle. However, residues 297 to 362 of the C-terminal arm had to be remodeled to accommodate the higher curvature of the T=1 particle. The loops, before and after the C-terminal short helix, were shown to provide the hinges that allowed curvature variation in the particle shell. The meeting densities seen at the threefold axes in the T=1 particle were consistent with the triple-helix interlinking contact at the local threefold axes in the T=7 structure.
Figures


Similar articles
-
New Structural Insights into the Genome and Minor Capsid Proteins of BK Polyomavirus using Cryo-Electron Microscopy.Structure. 2016 Apr 5;24(4):528-536. doi: 10.1016/j.str.2016.02.008. Epub 2016 Mar 17. Structure. 2016. PMID: 26996963 Free PMC article.
-
Characterization of self-assembled virus-like particles of human polyomavirus BK generated by recombinant baculoviruses.Virology. 2003 Jun 20;311(1):115-24. doi: 10.1016/s0042-6822(03)00141-7. Virology. 2003. PMID: 12832209
-
The refined structure of Nudaurelia capensis omega virus reveals control elements for a T = 4 capsid maturation.Virology. 2004 Jan 5;318(1):192-203. doi: 10.1016/j.virol.2003.08.045. Virology. 2004. PMID: 14972547
-
Host range analysis of simian virus 40, BK virus and chimaeric SV40/BKV: relative expression of large T-antigen and Vp1 in infected and transformed cells.Dev Biol Stand. 1998;94:191-205. Dev Biol Stand. 1998. PMID: 9776240 Review.
-
Recombinant virus like particles as drug delivery system.Curr Pharm Biotechnol. 2005 Feb;6(1):49-55. doi: 10.2174/1389201053167202. Curr Pharm Biotechnol. 2005. PMID: 15727555 Review.
Cited by
-
Leflunomide inhibition of BK virus replication in renal tubular epithelial cells.J Virol. 2010 Feb;84(4):2150-6. doi: 10.1128/JVI.01737-09. Epub 2009 Dec 2. J Virol. 2010. PMID: 19955306 Free PMC article.
-
Importance of calcium-binding site 2 in simian virus 40 infection.J Virol. 2007 Jun;81(11):6099-105. doi: 10.1128/JVI.02195-06. Epub 2007 Mar 14. J Virol. 2007. PMID: 17360742 Free PMC article.
-
The Symmetry of Viral Sialic Acid Binding Sites-Implications for Antiviral Strategies.Viruses. 2019 Oct 14;11(10):947. doi: 10.3390/v11100947. Viruses. 2019. PMID: 31615155 Free PMC article. Review.
-
New Structural Insights into the Genome and Minor Capsid Proteins of BK Polyomavirus using Cryo-Electron Microscopy.Structure. 2016 Apr 5;24(4):528-536. doi: 10.1016/j.str.2016.02.008. Epub 2016 Mar 17. Structure. 2016. PMID: 26996963 Free PMC article.
-
Biology of the BKPyV: An Update.Viruses. 2017 Nov 3;9(11):327. doi: 10.3390/v9110327. Viruses. 2017. PMID: 29099746 Free PMC article. Review.
References
-
- Baker, T. S., and R. H. Cheng. 1996. A model-based approach for determining orientations of biological macromolecules imaged by cryoelectron microscopy. J. Struct. Biol. 116:120-130. - PubMed
-
- Casini, G. L., D. Graham, D. Heine, R. L. Garcea, and D. T. Wu. 2004. In vitro papillomavirus capsid assembly analyzed by light scattering. Virology 325:320-327. - PubMed
-
- Chang, D., C. Y. Fung, W. C. Ou, P. C. Chao, S. Y. Li, M. Wang, Y. L. Huang, T. Y. Tzeng, and R. T. Tsai. 1997. Self-assembly of the JC virus major capsid protein, VP1, expressed in insect cells. J. Gen. Virol. 78:1435-1439. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

