The epsilon-subunit of mitochondrial ATP synthase is required for normal spindle orientation during the Drosophila embryonic divisions
- PMID: 15834145
- PMCID: PMC1450411
- DOI: 10.1534/genetics.104.037648
The epsilon-subunit of mitochondrial ATP synthase is required for normal spindle orientation during the Drosophila embryonic divisions
Abstract
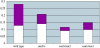
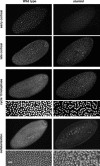
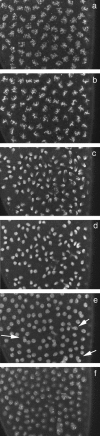
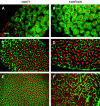
We describe the maternal-effect and zygotic phenotypes of null mutations in the Drosophila gene for the epsilon-subunit of mitochondrial ATP synthase, stunted (sun). Loss of zygotic sun expression leads to a dramatic delay in the growth rate of first instar larvae and ultimately death. Embryos lacking maternally supplied sun (sun embryos) have a sixfold reduction in ATP synthase activity. Cellular analysis of sun embryos shows defects only after the nuclei have migrated to the cortex. During the cortical divisions the actin-based metaphase and cellularization furrows do not form properly, and the nuclei show abnormal spacing and division failures. The most striking abnormality is that nuclei and spindles form lines and clusters, instead of adopting a regular spacing. This is reflected in a failure to properly position neighboring nonsister centrosomes during the telophase-to-interphase transition of the cortical divisions. Our study is consistent with a role for Sun in mitochondrial ATP synthesis and suggests that reduced ATP levels selectively affect molecular motors. As Sun has been identified as the ligand for the Methuselah receptor that regulates aging, Sun may function both within and outside mitochondria.
Figures


Similar articles
-
Spindle assembly and mitosis without centrosomes in parthenogenetic Sciara embryos.J Cell Biol. 1998 Jun 15;141(6):1383-91. doi: 10.1083/jcb.141.6.1383. J Cell Biol. 1998. PMID: 9628894 Free PMC article.
-
The 95F unconventional myosin is required for proper organization of the Drosophila syncytial blastoderm.J Cell Biol. 1995 Jun;129(6):1575-88. doi: 10.1083/jcb.129.6.1575. J Cell Biol. 1995. PMID: 7790355 Free PMC article.
-
Nuclear-fallout, a Drosophila protein that cycles from the cytoplasm to the centrosomes, regulates cortical microfilament organization.Development. 1998 Apr;125(7):1295-303. doi: 10.1242/dev.125.7.1295. Development. 1998. PMID: 9477328
-
Regulatory mechanisms of proton-translocating F(O)F (1)-ATP synthase.Results Probl Cell Differ. 2008;45:279-308. doi: 10.1007/400_2007_043. Results Probl Cell Differ. 2008. PMID: 18026702 Review.
-
Energetic signalling in the control of mitochondrial F1F0 ATP synthase activity in health and disease.Int J Biochem Cell Biol. 2008;40(12):2698-701. doi: 10.1016/j.biocel.2008.06.013. Epub 2008 Jul 30. Int J Biochem Cell Biol. 2008. PMID: 18707016 Review.
Cited by
-
Integrating animal development: How hormones and metabolism regulate developmental transitions and brain formation.Dev Biol. 2021 Jul;475:256-264. doi: 10.1016/j.ydbio.2021.01.016. Epub 2021 Feb 4. Dev Biol. 2021. PMID: 33549549 Free PMC article. Review.
-
ATP Synthase Subunit Epsilon Overexpression Promotes Metastasis by Modulating AMPK Signaling to Induce Epithelial-to-Mesenchymal Transition and Is a Poor Prognostic Marker in Colorectal Cancer Patients.J Clin Med. 2019 Jul 21;8(7):1070. doi: 10.3390/jcm8071070. J Clin Med. 2019. PMID: 31330880 Free PMC article.
-
SPD-3 is required for spindle alignment in Caenorhabditis elegans embryos and localizes to mitochondria.Genetics. 2007 Nov;177(3):1609-20. doi: 10.1534/genetics.107.078386. Epub 2007 Oct 18. Genetics. 2007. PMID: 17947426 Free PMC article.
-
Characterization of Drosophila ATPsynC mutants as a new model of mitochondrial ATP synthase disorders.PLoS One. 2018 Aug 10;13(8):e0201811. doi: 10.1371/journal.pone.0201811. eCollection 2018. PLoS One. 2018. PMID: 30096161 Free PMC article.
-
Combined positive effect of oocyte extracts and brilliant cresyl blue stained recipient cytoplasts on epigenetic reprogramming and gene expression in buffalo nuclear transfer embryos.Cytotechnology. 2017 Apr;69(2):289-305. doi: 10.1007/s10616-016-0057-0. Epub 2017 Jan 9. Cytotechnology. 2017. PMID: 28070808 Free PMC article.
References
-
- Ashburner, M., 1989 Drosophila: A Laboratory Handbook, pp. 345–346. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY.
-
- Bowman, B. J., S. E. Mainzer, K. E. Allen and C. W. Slayman, 1978. Effects of inhibitors on the plasma membrane and mitochondrial adenosine triphosphatases of Neurospora crassa. Biochim. Biophys. Acta 512: 13–28. - PubMed
-
- Boyer, P. D., 1997. The ATP synthase—a splendid molecular machine. Annu. Rev. Biochem. 66: 717–749. - PubMed
-
- Britton, J. S., and B. A. Edgar, 1998. Environmental control of the cell cycle in Drosophila: nutrition activates mitotic and endoreplicative cells by distinct mechanisms. Development 125: 2149–2158. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

