Hepatocyte growth factor is a preferred in vitro substrate for human hepsin, a membrane-anchored serine protease implicated in prostate and ovarian cancers
- PMID: 15839837
- PMCID: PMC1184568
- DOI: 10.1042/BJ20041955
Hepatocyte growth factor is a preferred in vitro substrate for human hepsin, a membrane-anchored serine protease implicated in prostate and ovarian cancers
Abstract
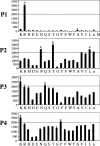
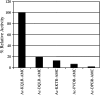
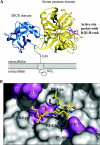
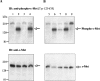
Hepsin is a membrane-anchored, trypsin-like serine protease with prominent expression in the human liver and tumours of the prostate and ovaries. To better understand the biological functions of hepsin, we identified macromolecular substrates employing a tetrapeptide PS-SCL (positional scanning-synthetic combinatorial library) screen that rapidly determines the P1-P4 substrate specificity. Hepsin exhibited strong preference at the P1 position for arginine over lysine, and favoured threonine, leucine or asparagine at the P2, glutamine or lysine at the P3, and proline or lysine at the P4 position. The relative activity of hepsin toward individual AMC (7-amino-4-methylcoumarin)-tetrapeptides was generally consistent with the overall peptide profiling results derived from the PC-SCL screen. The most active tetrapeptide substrate Ac (acetyl)-KQLR-AMC matched with the activation cleavage site of the hepatocyte growth factor precursor sc-HGF (single-chain HGF), KQLR downward arrowVVNG (where downward arrow denotes the cleavage site), as identified by a database analysis of trypsin-like precursors. X-ray crystallographic studies with KQLR chloromethylketone showed that the KQLR peptide fits well into the substrate-binding cleft of hepsin. This hepsin-processed HGF induced c-Met receptor tyrosine phosphorylation in SKOV-3 ovarian cancer cells, indicating that the hepsin-cleaved HGF is biologically active. Activation cleavage site mutants of sc-HGF with predicted non-preferred sequences, DPGR downward arrowVVNG or KQLQ downward arrowVVNG, were not processed, illustrating that the P4-P1 residues can be important determinants for substrate specificity. In addition to finding macromolecular hepsin substrates, the extracellular inhibitors of the HGF activator, HAI-1 and HAI-2, were potent inhibitors of hepsin activity (IC50 4+/-0.2 nM and 12+/-0.5 nM respectively). Together, our findings suggest that the HGF precursor is a potential in vivo substrate for hepsin in tumours, where hepsin expression is dysregulated and may influence tumorigenesis through inappropriate activation and/or regulation of HGF receptor (c-Met) functions.
Figures

References
-
- Leytus S. P., Loeb K. R., Hagen F. S., Kurachi K., Davie E. W. A novel trypsin-like serine protease (hepsin) with a putative transmembrane domain expressed by human liver and hepatoma cells. Biochemistry. 1988;27:1067–1074. - PubMed
-
- Kurachi K., Torres-Rosado A., Tsuji A. Hepsin. Methods Enzymol. 1994;244:100–114. - PubMed
-
- Netzel-Arnett S., Hooper J. D., Szabo R., Madison E. L., Quigley J. P., Bugge T. H., Antalis T. M. Membrane anchored serine proteases: a rapidly expanding group of cell surface proteolytic enzymes with potential roles in cancer. Cancer Metastasis Rev. 2003;22:237–258. - PubMed
-
- Wu Q. Type II transmembrane serine proteases. Curr. Opin. Dev. Biol. 2003;54:167–206. - PubMed
-
- Kazama Y., Hamamoto T., Foster D. C., Kisiel W. Hepsin, a putative membrane-associated serine protease, activates human factor VII and initiates a pathway of blood coagulation on the cell surface leading to thrombin formation. J. Biol. Chem. 1995;270:66–72. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

