Solving the protein sequence metric problem
- PMID: 15851683
- PMCID: PMC1088356
- DOI: 10.1073/pnas.0408677102
Solving the protein sequence metric problem
Abstract
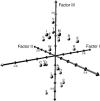
Biological sequences are composed of long strings of alphabetic letters rather than arrays of numerical values. Lack of a natural underlying metric for comparing such alphabetic data significantly inhibits sophisticated statistical analyses of sequences, modeling structural and functional aspects of proteins, and related problems. Herein, we use multivariate statistical analyses on almost 500 amino acid attributes to produce a small set of highly interpretable numeric patterns of amino acid variability. These high-dimensional attribute data are summarized by five multidimensional patterns of attribute covariation that reflect polarity, secondary structure, molecular volume, codon diversity, and electrostatic charge. Numerical scores for each amino acid then transform amino acid sequences for statistical analyses. Relationships between transformed data and amino acid substitution matrices show significant associations for polarity and codon diversity scores. Transformed alphabetic data are used in analysis of variance and discriminant analysis to study DNA binding in the basic helix-loop-helix proteins. The transformed scores offer a general solution for analyzing a wide variety of sequence analysis problems.
Figures

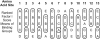
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

