Implications of the serine protease HtrA1 in amyloid precursor protein processing
- PMID: 15855271
- PMCID: PMC1087941
- DOI: 10.1073/pnas.0501823102
Implications of the serine protease HtrA1 in amyloid precursor protein processing
Abstract
The defining features of the widely conserved HtrA (high temperature requirement) family of serine proteases are the combination of a catalytic protease domain with one or more C-terminal PDZ domains and reversible zymogen activation. Even though HtrAs have previously been implicated in protein quality control and various diseases, including cancer, arthritis, and neuromuscular disorder, the biology of the human family members is not well understood. Our data suggest that HtrA1 is directly involved in the beta-amyloid pathway as it degrades various fragments of amyloid precursor protein while an HtrA1 inhibitor causes accumulation of Abeta in astrocyte cell culture supernatants. Furthermore, HtrA1 colocalizes with beta-amyloid deposits in human brain samples. Potential implications in Alzheimer's disease are discussed.
Figures

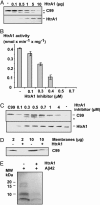


References
-
- Clausen, T., Southan, C. & Ehrmann, M. (2002) Mol. Cell 10, 443-455. - PubMed
-
- Ehrmann, M. & Clausen, T. (2004) Annu. Rev. Genet. 38, 709-724. - PubMed
-
- Spiess, C., Beil, A. & Ehrmann, M. (1999) Cell 97, 339-347. - PubMed
-
- Krojer, T., Garrido-Franco, M., Huber, R., Ehrmann, M. & Clausen, T. (2002) Nature 416, 455-459. - PubMed
-
- Walsh, N. P., Alba, B. M., Bose, B., Gross, C. A. & Sauer, R. T. (2003) Cell 113, 61-71. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

