Imidazole antibiotics inhibit the nitric oxide dioxygenase function of microbial flavohemoglobin
- PMID: 15855504
- PMCID: PMC1087630
- DOI: 10.1128/AAC.49.5.1837-1843.2005
Imidazole antibiotics inhibit the nitric oxide dioxygenase function of microbial flavohemoglobin
Abstract
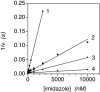
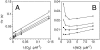
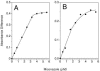
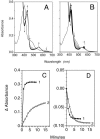
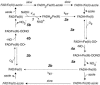
Flavohemoglobins metabolize nitric oxide (NO) to nitrate and protect bacteria and fungi from NO-mediated damage, growth inhibition, and killing by NO-releasing immune cells. Antimicrobial imidazoles were tested for their ability to coordinate flavohemoglobin and inhibit its NO dioxygenase (NOD) function. Miconazole, econazole, clotrimazole, and ketoconazole inhibited the NOD activity of Escherichia coli flavohemoglobin with apparent K(i) values of 80, 550, 1,300, and 5,000 nM, respectively. Saccharomyces cerevisiae, Candida albicans, and Alcaligenes eutrophus enzymes exhibited similar sensitivities to imidazoles. Imidazoles coordinated the heme iron atom, impaired ferric heme reduction, produced uncompetitive inhibition with respect to O(2) and NO, and inhibited NO metabolism by yeasts and bacteria. Nevertheless, these imidazoles were not sufficiently selective to fully mimic the NO-dependent growth stasis seen with NOD-deficient mutants. The results demonstrate a mechanism for NOD inhibition by imidazoles and suggest a target for imidazole engineering.
Figures

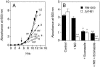
Similar articles
-
Catalytic Differences between Flavohemoglobins of Giardia intestinalis and E. coli.Pathogens. 2024 Jun 6;13(6):480. doi: 10.3390/pathogens13060480. Pathogens. 2024. PMID: 38921778 Free PMC article.
-
Structural studies on flavohemoglobins.Methods Enzymol. 2008;436:187-202. doi: 10.1016/S0076-6879(08)36010-8. Methods Enzymol. 2008. PMID: 18237633
-
Bacterial flavohemoglobin: a molecular tool to probe mammalian nitric oxide biology.Biotechniques. 2011 Jan;50(1):41-5. doi: 10.2144/000113586. Biotechniques. 2011. PMID: 21231921 Free PMC article.
-
Nitric oxide dioxygenase function and mechanism of flavohemoglobin, hemoglobin, myoglobin and their associated reductases.J Inorg Biochem. 2005 Jan;99(1):247-66. doi: 10.1016/j.jinorgbio.2004.10.003. J Inorg Biochem. 2005. PMID: 15598505 Review.
-
A survey of methods for the purification of microbial flavohemoglobins.Methods Enzymol. 2008;436:169-86. doi: 10.1016/S0076-6879(08)36009-1. Methods Enzymol. 2008. PMID: 18237632 Review.
Cited by
-
Synthesis, characterization, in silico molecular docking, and antibacterial activities of some new nitrogen-heterocyclic analogues based on a p-phenolic unit.RSC Adv. 2022 Apr 26;12(20):12607-12621. doi: 10.1039/d2ra01794f. eCollection 2022 Apr 22. RSC Adv. 2022. PMID: 35496342 Free PMC article.
-
Cobalamin in inflammation III - glutathionylcobalamin and methylcobalamin/adenosylcobalamin coenzymes: the sword in the stone? How cobalamin may directly regulate the nitric oxide synthases.J Nutr Environ Med. 2007 Sep;16(3-4):212-226. doi: 10.1080/13590840701791863. Epub 2008 Jan 10. J Nutr Environ Med. 2007. PMID: 18923642 Free PMC article.
-
Imidazoles and Quaternary Ammonium Compounds as Effective Therapies against (Multidrug-Resistant) Bacterial Wound Infections.Antibiotics (Basel). 2024 Oct 10;13(10):949. doi: 10.3390/antibiotics13100949. Antibiotics (Basel). 2024. PMID: 39452215 Free PMC article.
-
Comparative genome analysis provides insights into the evolution and adaptation of Pseudomonas syringae pv. aesculi on Aesculus hippocastanum.PLoS One. 2010 Apr 19;5(4):e10224. doi: 10.1371/journal.pone.0010224. PLoS One. 2010. PMID: 20419105 Free PMC article.
-
Allostery in the nitric oxide dioxygenase mechanism of flavohemoglobin.J Biol Chem. 2021 Jan-Jun;296:100186. doi: 10.1074/jbc.RA120.016637. Epub 2020 Dec 17. J Biol Chem. 2021. PMID: 33310705 Free PMC article.
References
-
- Cornelissen, F., and H. Van den Bossche. 1983. Synergism of the antimicrobial agents miconazole, bactracin and polymyxin B. Chemotherapy 29:419-427. - PubMed
-
- Crawford, M. J., and D. E. Goldberg. 1998. Regulation of the Salmonella typhimurium flavohemoglobin gene. A new pathway for bacterial gene expression in response to nitric oxide. J. Biol. Chem. 273:34028-34032. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

