Aldolases a and C are ribonucleolytic components of a neuronal complex that regulates the stability of the light-neurofilament mRNA
- PMID: 15858061
- PMCID: PMC6725117
- DOI: 10.1523/JNEUROSCI.0885-05.2005
Aldolases a and C are ribonucleolytic components of a neuronal complex that regulates the stability of the light-neurofilament mRNA
Erratum in
- J Neurosci. 2006 May 10;26(19):5276
Abstract
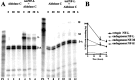
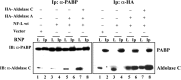
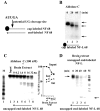
A 68 nucleotide segment of the light neurofilament (NF-L) mRNA, spanning the translation termination signal, participates in regulating the stability of the transcript in vivo. Aldolases A and C, but not B, interact specifically with this segment of the transcript in vitro. Aldolases A and C are glycolytic enzymes expressed in neural cells, and their mRNA binding activity represents a novel function of these isozymes. This unsuspected new activity was first uncovered by Northwestern blotting of a brainstem/spinal cord cDNA library. It was confirmed by two-dimensional fractionation of mouse brain cytosol followed by Northwestern hybridization and protein sequencing. Both neuronal aldolases interact specifically with the NF-L but not the heavy neurofilament mRNA, and their binding to the transcript excludes the poly(A)-binding protein (PABP) from the complex. Constitutive ectopic expression of aldolases A and C accelerates the decay of a neurofilament transgene (NF-L) driven by a tetracycline inducible system. In contrast, mutant transgenes lacking mRNA sequence for aldolase binding are stabilized. Our findings strongly suggest that aldolases A and C are regulatory components of a light neurofilament mRNA complex that modulates the stability of NF-L mRNA. This modulation likely involves endonucleolytic cleavage and a competing interaction with the PABP. Interactions of aldolases A and C in NF-L expression may be linked to regulatory pathways that maintain the highly asymmetrical form and function of large neurons.
Figures




Similar articles
-
Coregulation of light neurofilament mRNA by poly(A)-binding protein and aldolase C: implications for neurodegeneration.Brain Res. 2007 Mar 30;1139:15-28. doi: 10.1016/j.brainres.2006.12.092. Epub 2007 Jan 10. Brain Res. 2007. PMID: 17276415
-
Similar poly(C)-sensitive RNA-binding complexes regulate the stability of the heavy and light neurofilament mRNAs.Brain Res. 2000 Jun 9;867(1-2):265-79. doi: 10.1016/s0006-8993(00)02389-1. Brain Res. 2000. PMID: 10837825
-
3' untranslated region in a light neurofilament (NF-L) mRNA triggers aggregation of NF-L and mutant superoxide dismutase 1 proteins in neuronal cells.J Neurosci. 2004 Mar 17;24(11):2716-26. doi: 10.1523/JNEUROSCI.5689-03.2004. J Neurosci. 2004. PMID: 15028764 Free PMC article.
-
Stability determinants are localized to the 3'-untranslated region and 3'-coding region of the neurofilament light subunit mRNA using a tetracycline-inducible promoter.J Biol Chem. 1998 May 15;273(20):12650-4. doi: 10.1074/jbc.273.20.12650. J Biol Chem. 1998. PMID: 9575227
-
A novel RNA binding protein that interacts with NMDA R1 mRNA: regulation by ethanol.Eur J Neurosci. 2006 May;23(9):2339-50. doi: 10.1111/j.1460-9568.2006.04776.x. Eur J Neurosci. 2006. PMID: 16706842
Cited by
-
TMT-Based Proteomic Analysis of Plasma from Children with Rolandic Epilepsy.Dis Markers. 2020 Oct 7;2020:8840482. doi: 10.1155/2020/8840482. eCollection 2020. Dis Markers. 2020. PMID: 33082886 Free PMC article.
-
The role of mammalian poly(A)-binding proteins in co-ordinating mRNA turnover.Biochem Soc Trans. 2012 Aug;40(4):856-64. doi: 10.1042/BST20120100. Biochem Soc Trans. 2012. PMID: 22817748 Free PMC article. Review.
-
High content screening of cortical neurons identifies novel regulators of axon growth.Mol Cell Neurosci. 2010 May;44(1):43-54. doi: 10.1016/j.mcn.2010.02.002. Epub 2010 Feb 14. Mol Cell Neurosci. 2010. PMID: 20159039 Free PMC article.
-
Detailed expression pattern of aldolase C (Aldoc) in the cerebellum, retina and other areas of the CNS studied in Aldoc-Venus knock-in mice.PLoS One. 2014 Jan 27;9(1):e86679. doi: 10.1371/journal.pone.0086679. eCollection 2014. PLoS One. 2014. PMID: 24475166 Free PMC article.
-
Cellular Prion Protein (PrPc): Putative Interacting Partners and Consequences of the Interaction.Int J Mol Sci. 2020 Sep 25;21(19):7058. doi: 10.3390/ijms21197058. Int J Mol Sci. 2020. PMID: 32992764 Free PMC article. Review.
References
-
- Ahn AH, Dziennis S, Hawkes R, Herrup K (1994) The cloning of zebrin II reveals its identity with aldolase C. Development 120: 2081-2090. - PubMed
-
- Amrani N, Ganesan R, Kervestin S, Mangus DA, Ghosh S, Jacobson A (2004) A faux 3′-UTR promotes aberrant termination and triggers non-sense-mediated mRNA decay. Nature 432: 118-129. - PubMed
-
- Berardini TZ, Drygas-Williams M, Callard GV, Tolan DR (1997) Identification of neuronal isozyme specific residues by comparison of goldfish aldolase C to other aldolases. Comp Biochem Physiol 117A: 471-476. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
